Abstract
In order to assess the ecological and genetic effects of cutting, we compared two portions of Alnus trabeculosa population at Yuda (Iwate Prefecture, Japan): one that has been cut about 30 years ago and one that has remained uncut. These portions were compared in terms of the degree of sprouting, genetic variation and gene distribution using isozyme markers. First, we determined the multilocus genotype (MLG) of all ramets, then sorted them into individuals according to the distribution of the MLGs. The average (±SE) of largest distance between ramets in one individual was 2.1 (±0.18) m, which is consistent with the distance (2.0 (±0.20) m) obtained by tracing physical connections between ramets. We found no significant differences in genetic variation between the two portions, but there were significant differences in their degree of sprouting. Furthermore, there were striking differences in gene distribution: the cut portion showed greater clustering of individuals with identical genetic components, which may be due to regeneration in the gaps made by cutting, reflecting the location of the mother trees, and seed and pollen dispersal from them.
Similar content being viewed by others
Introduction
Alnus trabeculosa Hand.-Mazz. is a deciduous, broad-leaved tree, which grows only in open swampy places, and reaches a height of 5 m. The species is sparsely distributed in the warm- and intermediate-temperate zones of Honshu and Kyushu (Japan), with northernmost limits in Yuda (Iwate Prefecture). It is also found in the southeastern part of continental China (Kurata, 1971). Since its habitat is dwindling and human activities are having an adverse influence on it, populations of the species are declining, and it is now classed as ‘near threatened’ (Environment Agency of Japan, 2000).
This species has the capacity to sprout, as well as to reproduce sexually via seeds (Handa et al, 1996). In accordance with this, some individuals in the Yuda population were observed to form sprouting clonal stands, and the extent of each individual could not be determined by simple visual inspection.
Genetic variation and, especially, family structure can result from several evolutionary processes, including isolation in small patches, limited pollen or seed dispersal, selection and human influences (Sokal and Menozzi, 1982). As a tool for analyzing family structure, spatial autocorrelation has been used in several studies of forest tree populations. For example, differences in genetic structure between populations with different age classes have been detected (Hamrick et al, 1993; Epperson and Alvarez-Buylla, 1997; Parker et al, 2001); genetic structure found in younger classes disappeared in older classes as the population was thinned. Strong structuring has also been detected using spatial autocorrelation in a population in which vegetative reproduction and limited dispersal of pollen and seeds were observed (Shapcott, 1995). It has also revealed striking differences in genetic structure between populations with differing histories of anthropogenic disturbance (Knowles et al, 1992; Takahashi et al, 2000).
Assessing subtle differences among evolutionary processes using spatial autocorrelation is unrealistic. Nevertheless, investigating family structure can help to describe the main evolutionary forces affecting the population under study (Sokal and Menozzi, 1982), and provide fundamental information needed for selecting natural populations for conservation, or breeding programs. Family structure should also be taken into account in order to maximize diversity, and to avoid making erroneous estimates of population diversity (Epperson, 1989).
Our group has previously detected the existence of family structure in the Juo population of A. trabeculosa in Ibaraki, central Japan, using spatial autocorrelation (Miyamoto et al, 2000, 2002). We concluded that for both in situ and ex situ conservation of this species, it is important to take into consideration the existence of family structure. We also suggested that the species' mode of seed dispersal and asexual propagation may be involved in the formation of family structure.
According to the owner of this site, part of the Yuda population was cut down about 30 years ago (pers. comm.). There now appear to be differences between the cut and uncut portions. The height and diameter of the trees in the harvested portion are smaller than those in the uncut portion. However, it is not clear whether there are ecological and/or genetic differences between these two A. trabeculosa portions.
Therefore, in order to assess the influence of cutting, we compared two portions of the Yuda population. First, we obtained information about multilocus genotypes (Okuizumi, 1993) for all the ramets, using isozyme markers. When ramets with identical MLGs are clustered, they are assumed to belong to a single individual. In order to identify individuals which are composed of one or more ramets with identical MLGs, a statistical method was used. Finally, we evaluated the degree of sprouting (in terms of number of sprouts per individual and the largest distance between ramets in one individual) as well as the genetic variation in the two portions with different cutting histories, and compared the pattern of gene distribution between the two portions using spatial autocorrelation analysis.
Materials and methods
Studied population
We studied the population of A. trabeculosa at Ecchu-hata, Yuda, in Iwate Prefecture (285 m a.s.l.; 39°17′N, 140°43′E; average temperature=9.5°C; warmth index (Σ(Tm−5), when Tm is above 5, Tm: monthly mean temperature)=78.9°C per month; coldness index (−Σ(5−Tm), when Tm is below 5) =−23.2°C per month; precipitation=1605 mm per year). This population, the northernmost population of the species in Japan, is located along a small river, which runs from west to east. The population is surrounded by a paddy field to the north, an artificial forest of Cryptomeria japonica (Japanese Cedar) to the south and a field to the west, where soil removed during the construction of the Akita Highway was dumped. This population is divided into two portions (western and eastern) that are 50 m apart. The western portion has many small trees and a few large ones, while, in contrast, the eastern portion is composed almost entirely of large trees. Trees in the western portion (portion A) are generally much smaller in height and diameter than those in the eastern portion (portion B), and were assumed to be immature. Judging from the distribution of genotypes among the ramets (see below), some plants in the investigated population seem to form sprouting, clonal stands. We defined genetically identical organisms derived from sexual propagation as ‘individuals’, and sprouts formed by asexual propagation belonging to a single individual as ‘ramets’. We also surveyed the height and diameter at breast height (DBH) of all ramets taller than 1 m (532 ramets) and mapped them (Figure 1).
Isozyme analysis
Experimental samples (winter buds) were collected from all mapped ramets in the winter of 1998. Samples were stored at −80°C until the enzymes were extracted. Isozyme analysis was based on the protocol described by Tsumura et al (1990). In A. trabeculosa, 12 putative polymorphic isozyme loci have already been reported, encoded by eight enzyme systems (Miyamoto et al, 2000): Aap (alanine aminopeptidase; AAP, E.C. 3.4.11.1), Aco (aconitase: ACO, E.C. 4.2.1.3), Amy-2 (amylase; AMY, E.C. 3.2.1), Dia-1, Dia-2, Dia-3 (diaphorase; DIA, E.C. 1.6.4.3), Got-1, Got-2 (glutamate oxaloacetate transaminase; GOT, E.C. 2.6.1.1), Lap (leucine aminopeptidase; LAP, E.C. 3.4.11.1), Pgi-1, Pgi-2 (phosphoglucoisomerase; PGI, E.C. 5.3.1.9) and Shd (shikimate dehydrogenase; ShDH, E.C. 1.1.1.25). These loci were used for the analysis.
Identification of individuals by the MLG of ramets
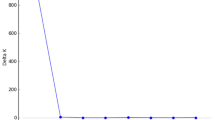
Information about the isozyme genotype at all 12 loci (multilocus genotype, MLG; Okuizumi, 1993) was obtained for all the ramets. When ramets with identical MLGs are clustered, they are assumed to belong to a single individual. In order to identify individuals, which are composed of ramets with identical MLGs located continuously, a statistical method was used. We made 20 distance classes of 1 m intervals (0–1, 1–2 m and so on). The number of joins of ramets with identical MLGs was counted in each distance class. For each MLG, the standard normal deviate (SND) of the observed number of joins from the expected number of joins (Sokal and Oden, 1978) was calculated using the formula:
The SND is assumed to be normally distributed, so it has critical values of ±1.96 at the 5% level of significance. If ramets with identical MLGs are located closer to each other, than would be expected according to random chance, the SND is expected to decrease as the distance class increases. Therefore, we examined the largest distance classes where SND scored significant values for each MLG and adopted its value to define individual boundaries. We then sorted the ramets with identical MLGs into individuals according to these values. The SND values were calculated using the software developed by Takahashi (2001).
The spread and type of individuals
The spread of each individual was defined as the distance between its two most widely spaced ramets. We investigated the spread of each individual identified in the way stated above. Using t-tests, we then compared the differences in the spread between individuals belonging to the two portions. Moreover, we classified the individuals into two types: those with only one ramet (type 1) and those with two or more ramets (type 2). We also calculated the percentage of each type, the average number of ramets per individual of type 2, and evaluated the significance of the difference in the number of ramets per individual of type 2 between the two portions using a t-test.
Physical analysis by root excavation
We dug down to the roots of each individual at two points and traced physical connections between ramets. In addition, we measured the largest distance between connected ramets belonging to each individual.
Genetic variation
To estimate genetic variation, the following genetic parameters (Hamrick and Godt, 1989) were calculated for all individuals of portions A and B: the percentage of polymorphic loci (Pl; we regarded the loci where frequency of the most frequent allele is less than 0.99 as polymorphic loci), the mean number of alleles per locus (Na), the effective number of alleles per locus (Ne), the observed heterozygosity (Ho) and the expected heterozygosity (He). The Ne values were derived from 1/Σpi2,' (Kimura and Crow, 1964) and He values by averaging he over 12 loci, where he=2n(1−Σpi2)/(2n−1) (Nei, 1987), pi is the allele frequency of the ith allele at a given locus and n is the sample size.
Gene distribution analysis in each portion
Spatial genetic structure within the two portions was assessed by estimating coancestry (Cockerham, 1969) between all possible pairs of individuals at different interplant distances. For each individual containing more than two ramets, we assumed that one ramet was at its center of gravity.
In order to estimate coancestry, genotypes for individuals were first expressed as allele frequencies of 1.0, 0.5 or 0.0, depending on whether their genotype included 2, 1 or 0 copies of the respective alleles (Heywood, 1991). Estimates of coancestry (ρ̂ij) were then calculated as

where pi and pj are the frequencies of same alleles at a locus for individuals i and j, p̄ is the mean frequency for that allele and k=n(n−1)/2, which gives the number of possible pairs between n individuals located in each distance class (Loiselle et al, 1995). The second term adjusts for bias associated with a finite sample size, which causes the estimate of coancestry to be zero for a population in Hardy–Weinberg equilibrium. The correlation between pairs of plants in their frequencies of same alleles at a given locus (ρ̂ij) in an outbreeding population (FIS=0) has expected values of 0.125 and 0.0625 for half-sibs and first cousins, respectively. A combined multilocus estimate of coancestry can be obtained by weighting the values for each locus by its polymorphic index, Σpi(1−pi). Weights for individual loci were also adjusted for differences in sample size due to missing genotypes (Loiselle et al, 1995). Multilocus coancestry estimates have a sound foundation in population genetics theory and provide a natural means of summarizing data over alleles at a locus and over loci to provide a more powerful test for spatial genetic structure (Heywood, 1991). Therefore, multilocus estimates of coancestry were calculated at 5 m interplant-distance increments for each subpopulation. Tests of significance for estimated values of ρ̂ij were performed using randomization procedures to generate populations under the null hypothesis of no spatial structure. In each subpopulation, intact multilocus genotypes were randomly drawn, with replacement, from the sampled data and assigned to points occupied by plants. New ρ̂ij values, were then calculated. This randomization procedure was repeated 399 times for each subpopulation, giving (together with the originally sampled data) 400 ρ̂ij values, from which 95 and 99% confidence intervals were constructed. The procedure tests the significance of the correlation in multilocus genotypes at each distance interval (Slatkin and Arter, 1991). We used software developed by Nason (1997), (1998) for the autocorrelation analyses of coancestry.
Results
Size analysis in each portion
The means and standard deviations of the height and DBH values of ramets in the Yuda population were 5.7 ±2.7 m and 8.1±4.2 cm, respectively. The corresponding values were 4.3±1.7 m and 6.2±2.9 cm, respectively, for ramets in portion A, compared to 8.1±2.4 m and 11.6±4.1 cm in portion B (Table 1;; Figure 2). Thus, ramets in portion B were approximately twice as tall and twice as wide as those in portion A, and differences between the ramets in the two portions in both height and diameter at breast height were significant (p<0.01 and p<0.01, respectively).
Isozyme analysis
Variation was detected at five of the 12 putative isozyme loci tested (Aap, Amy-2, Got-2, Lap and Shd).
Identification of individuals by the MLGs of ramets
Individuals were identified based on the results of the MLG analysis and evaluation of the distance classes for which the MLGs had significant SND values. A total of 532 ramets were sorted into 225 individuals by 57 MLGs. In all, 134 and 91 individuals were recognized from portions A and B, respectively.
The spread and type of individuals
The proportion of type 2 individuals was very similar in the two portions; 51% in portion A and 48% in portion B (Table 1). However, the average number of ramets per type 2 individual was higher in portion A (283/68 or 4.2) than in portion B (136/44 or 3.1), and there was a significant difference between these numbers according to t-tests (p<0.05). The largest individual was 10.8 m wide in portion A. The most frequently observed spread class was 1.0–2.0 m, and the average spread of these individuals (±SE) was 2.1±0.18 m (2.3±0.27 and 1.6±0.20 m in portions A and B, respectively; Figure 3). There was a significant difference between these values according to t-tests (p<0.05).
Physical analysis by root excavation
Physical connections between ramets of two individuals were consistent with the identification based on the MLG analysis. Two individuals showed that ramets were connected within the range of 2.0±0.20 m. This was almost the same as the mean value derived from the MGC analysis (2.1±0.18 m).
Genetic variation
No significant differences were found between the two portions in genetic diversity parameters (Table 2).
Gene distribution analysis in each portion
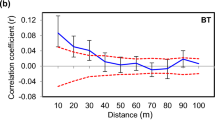
In portion B, coancestry values were within the range of 99% confidence limits in all of the distance classes, that is, coancestry values were not significantly different from zero. In contrast, the coancestry coefficients in portion A were higher than the expected range in the first two distance classes, but fell to within the expected interval in the third distance class (upper limit, 15 m). These data indicate that individuals within 10 m of each other tend to be genetically similar in portion A (Figure 4).
Discussion
Influence of cutting on sprouting
A. trabeculosa has previously been reported to produce sprouts from its roots (Handa et al, 1996). The distribution of trunks we found with the same MLGs strongly corroborates this report. We estimated the influence of cutting in terms of the spread of area spanned by individuals and the number of ramets belonging to each individual. The maximum spread of an individual clone is about 10 m, and the average range of individuals including more than two ramets is about 2.1 m. This value is consistent with the range (2.0 m) obtained by physically tracing connections through root excavation. Previously reported ranges of individuals or clumps of ramets include 2–4 m in Fagus japonica (Kitamura et al, 1992), 4 m in Quercus chrysolepis (Montalvo et al, 1997), 8 m in Fagus grandifolia (Jones and Raynal, 1987), and areas of 50 m2 in Thujopsis dolabrata (Hashimoto et al, 1999) and up to 43 ha in Populus tremuloides (Kemperman and Barnes, 1976). Thus, the A. trabeculosa range found in this study was similar to those of F. japonica and Q. chrysolepis.
Comparison of the height and DBH of trees in the two portions showed that ramets in portion B, which had not been cut, were about twice as large as those in portion A. which has been cut. The proportion of type 2 individuals, that is, those with two or more ramets, was similar in portions A and B, but the average number of ramets per type 2 individual was significantly higher in portion A. Two factors may have contributed to this difference. Firstly, type 2 individuals may become type 1 individuals, as some ramets die. Alternatively, since portion A is considered to be younger and immature, the stand in portion A may be in a transitional state towards that of portion B. Secondly, tree species are generally thought to increase by asexual propagation when the trunks die due to damage caused by fungi, or because they have fallen down or been blown down by the wind (Hallé et al, 1978; Watanabe, 1994). Portion A was cut down approximately 30 years ago; so it is presumed that the cutting might have induced sprouting in this species.
Influence of cutting on the genetic variation and gene distribution
No significant differences were found between the two portions in terms of genetic diversity parameters. However, the coancestry analyses revealed striking differences between the allele and genotype distributions in the two portions. Portion A showed higher degrees of clustering of individuals with identical genes than portion B, and the range of the clusters with similar genetic components appears to be up to 10 m. Since portion A was partly cut down in the past, it is now thought to be composed of large remnant trees together with small seedlings or sprouts from the trees that were cut down. There are many potential candidates for the mother trees of these seedlings, but it is highly likely that they were trees that were cut and/or remnant trees. In contrast, in portion B, the mother trees may have been located in the other portion, and dispersed seeds by wind to portion B, or they may have dropped seeds that floated to the present portion B, which was a safe site for A. trabeculosa. So, we can speculate that the gene and genotype concentrations seen in portion A reflect the locations and limited pool of mother trees before and just after the cutting, while in portion B, clustering of genes and genotypes occurs to a much lower degree because the seeds came from more widely dispersed trees.
Miyamoto et al (2000) suggested that the mode of seed dispersal and asexual propagation in the Juo population of the species may be related to the formation of family structure. In addition, this study suggests that the cutting may have caused clustering of individuals with identical genes and thus created the family structure. This indicates that the uncut portion may be more suitable for conservation, because of its more random distribution of genes, than the portion that was cut in the past.
References
Cockerham CC (1969). Variance of gene frequencies. Evolution 23: 72–84.
Environment Agency of Japan (2000). Threatened Wildlife of Japan – Red Data Book, 2nd edn., Vol. 8, Vascular Plants. Japan Wildlife Research Center: Tokyo. 660pp. (in Japanese).
Epperson BK (1989). Spatial patterns of genetic variation within plant populations. In: Brown AHD, Clegg MT, Kahler AL, Weir BS (eds) Plant Population Genetics, Breeding and Genetic Resources, Sinauer Associates: Sunderland, MA. pp 229–253.
Epperson BK, Alvarez-Buylla ER (1997). Limited seed dispersal and genetic structure in life stages of Cecropia obtusifolia. Evolution 51: 275–282.
Hallé F, Oldeman RAA, Tomlinson PB (1978). Tropical Trees and Forests: An Architectural Analysis, Springer-Verlag: Berlin.
Hamrick JL, Godt MJW (1989). Allozyme diversity in plant species. In: Brown AHD, Clegg MT, Kahler AL, Weir BS (eds) Plant Population Genetics, Breeding and Genetic Resources, Sinauer Associates: Sunderland, MA. pp 43–63.
Hamrick JL, Murawski DA, Nason JD (1993). The influence of seed dispersal mechanisms on the genetic structure of tropical tree populations. Vegetation 107/108: 281–297.
Handa T, Senda M, Hoshi H, Nishimura K (1996) Alnus trabeculosa population at Juo in Ibaraki Prefecture (1). Trans Jpn For Soc 107: 145 (in Japanese).
Hashimoto R, Yamaguchi R, Satou N (1999). Studies on occurrence of layering clumps and regeneration pattern of Thujopsis dolabrata var. hondai by isozyme variations. J Jpn For Soc 81: 169–177.
Heywood JS (1991). Spatial analysis of genetic variation in plant populations. Ann Rev Ecol Syst 22: 335–355.
Jones RH, Raynal DJ (1987). Root sprouting in American beech: production, survival, and the effect of parent tree vigor. Can J For Res 17: 539–544.
Kemperman JA, Barnes BV (1976). Clone size in American aspens. Can J Bot 54: 2603–2607.
Kimura M, Crow JF (1964). The number of alleles that can be maintained in a finite population. Genetics 49: 725–738.
Kitamura K, Okuizumi H, Seiki T, Niiyama K, Shiraishi S (1992). Isozyme analysis of the mating system in natural populations of Fagus crenata and F. japonica. Jpn J Ecol 42: 61–69.
Knowles P, Perry DJ, Forster HA (1992). Spatial genetic structure in two tamarack (Larix laricina (Du Roi) K. Koch) populations with differing establishment histories. Evolution 46: 572–576.
Kurata S (1971). Illustrated Important Forest Trees of Japan, Chikyu shuppan: Tokyo. Vol. 3, 259pp (in Japanese).
Loiselle BA, Sork VL, Nason J, Graham C (1995). Spatial genetic structure of a tropical understory shrub Psychotria officinalis (Rubiaceae). Am Bot 82: 1420–1425.
Miyamoto N, Kuramoto N, Yamada H (2002). Differences in spatial autocorrelation between four sub-populations of Alnns trabeculosa Hand.-Mazz. (Betulaceae). Heredity 89: 273–279.
Miyamoto N, Yamamoto N, Hoshi H, Handa T (2000). Isozyme polymorphisms and family structure of Alnus trabeculosa Hand.-Mazz. (Betulaceae) population at Juo, Ibaraki Prefecture. J Jpn For Soc 82: 72–79 (in Japanese with English summary).
Montalvo AL, Conard SG, Conkle MT, Hodgskiss PD (1997). Population structure, genetic diversity, and clone formation in Quercus chrysolepis (Fagaceae). Am J Bot 84: 1553–1564.
Nason JD (1997). FijAnal. A computer program for the analysis of spatial autocorrelation. Version 2.1. http://www.nceas.ucsb.edu/papers/geneflow/software/.
Nason JD (1998). BS_fij. A computer program for the analysis of significance tests. Version 2.1. Free program distributed by the author over the internet from http://www.nceas.ucsb.edu/papers/geneflow/software/.
Nei M (1987). Molecular Evolutionary Genetics, Columbia University Press: New York.
Okuizumi H (1993). Clone analysis of collected sugi-cutting cultivars of the Kyusyu Region by the multilocus genotypes of twelve isozyme loci. J Jpn For Soc 75: 293–302.
Parker KC, Hamrick JL, Parker AJ, Nason JD (2001). Fine-scale genetic structure in Pinus clausa (Pinaceae) populations: effects of disturbance history. Heredity 87: 99–113.
Shapcott A (1995). The spatial genetic structure in natural populations of the Australian temperate rainforest tree Atherosperma moschatum. (Labill.) (Monimiaceae). Heredity 74: 28–38.
Slatkin M, Arter HE (1991). Spatial autocorrelation methods in population genetics. Am Nat 138: 449–517.
Sokal RR, Menozzi P (1982). Spatial autocorrelation of HLA frequencies in Europe support demic diffusion of early farmers. Am Nat 119: 1–17.
Sokal RR, Oden DL (1978). Spatial autocorrelation in biology. 1. Methodology. Biol J Linn Soc 10: 199–228.
Takahashi M (2001). A freeware program of spatial autocorrelation analysis for Windows 95 and 98. Delphi type, version 1.1.1. http://150.26.104.1/makotoHP/FrameE.htm.
Takahashi M, Mukouda M, Koono K (2000). Differences in genetic structure between two Japanese beech (Fagus crenata Blume) stands. Heredity 84: 103–115.
Tsumura Y, Tomaru N, Suyama Y, M Na'iem, Ohba K (1990). Laboratory manual of isozyme analysis. Bull Tsukuba Univ For 6: 63–95 (in Japanese).
Watanabe S (1994). Specia of Trees. University of Tokyo Press: Tokyo. 450 pp (in Japanese).
Acknowledgements
We thank Mr K Hayakawa and the committee of education in Yuda, Iwate Prefecture for offering access to the field site and for providing useful information, Mr M Ueta, Mr N Miura, Mr T Hasebe and Ms T Wakai for their help in collecting samples and Ms K Tanaka and Ms Y Miyata for their assistance in the laboratory.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Miyamoto, N., Karamoto, N. & Takahashi, M. Ecological and genetic effects of cutting in an Alnus trabeculosa Hand.-Mazz. (Betulaceae) population. Heredity 91, 331–336 (2003). https://doi.org/10.1038/sj.hdy.6800316
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800316
Keywords
This article is cited by
-
Fine-scale genetic structure within plots of Polygala reinii (Polygalaceae) having an ant-dispersal seed
Journal of Plant Research (2010)
-
A plant nutrition strategy for ex-situ conservation based on “Ecological Similarity”
Journal of Forestry Research (2008)