Abstract
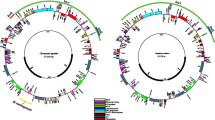
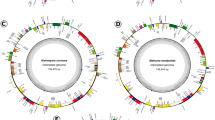
Discocactus and Melocactus are representatives of Core Cactoideae II BCT clade (Browningieae, Cereeae and Trichocereeae clade) and consist of 14 and 50 species, respectively. Both genera occur in South America, but Melocactus distribution extends through Central America and the West Indies. Although the Cactaceae family stands out for having highly rearranged plastomes, with particular gene organization, small or divergent inverted repeats (IRs) and IR loss, the plastome variation in the BCT clade is unknown. In this study, we assembled plastomes of Discocactus bahiensis and Melocactus ernestii and compared them with other available plastomes of Cactoideae (Carnegiea gigantea, and species of the genera Mammillaria and Rhipsalis) as well as with canonical plastomes of members of the order Caryophyllales (Portulaca oleracea and Spinacea oleracea). The plastomes of D. bahiensis (128,733-bp) and M. ernestii (130,703-bp) showed a quadripartite structure with 17 rearrangements when compared to those of P. oleracea and S. oleracea. Thus, twenty-eight genes were transferred from the LSC (large single copy) region to the IRs and 12 genes from the IRs to the LSC or SSC (small single copy) regions. Twenty-seven syntenic blocks were identified, evidencing a large number of rearrangements within the subfamily Cactoideae. These rearrangements could not be explained by the absence of IR regions nor the number of repeats. By comparing plastomes of D. bahiensis and M. ernestii, we propose new regions for phylogenetic inferences for the subfamily Cactoideae, for Discocactus and Melocactus, and for related groups.


Similar content being viewed by others
References
Anderson EF (2001) The cactus family. Timber Press, Portland
Arakaki M, Christin P-A, Nyffeler R, Lendel A, Eggli U, Ogburn RM, Spriggs E, Moore MJ, Edwards EJ (2011) Contemporaneous and recent radiations of the world’s major succulent plant lineages. Proc Natl Acad Sci 108:8379–8384. https://doi.org/10.1073/pnas.1100628108
Barrett CF, Davis JI (2012) The plastid genome of the mycoheterotrophic Corallorhiza striata (Orchidaceae) is in the relatively early stages of degradation. Am J Bot 99:1513–1523. https://doi.org/10.3732/ajb.1200256
Barrett CF, Freudenstein JV, Li J, Mayfield-Jones DR, Perez L, Pires JC, Santos C (2014) Investigating the path of plastid genome degradation in an early-transitional clade of heterotrophic orchids, and implications for heterotrophic angiosperms. Mol Biol Evol 31:3095–3112. https://doi.org/10.1093/molbev/msu252
Barrett CF, Baker WJ, Comer JR et al (2015) Plastid genomes reveal support for deep phylogenetic relationships and extensive rate variation among palms and other commelinid monocots. New Phytol 209:855–870. https://doi.org/10.1111/nph.13617
Barthlott W, Hunt DR (1993) Cactaceae. In: Kubitzki K, Rohwer JG, Bittrich V (eds) Flowering plants dicotyledons. Springer, Berlin Heidelberg, Berlin, pp 161–197
Blazier JC, Jansen RK, Mower JP, Govindu M, Zhang J, Weng M-L, Ruhlman TA (2016) Variable presence of the inverted repeat and plastome stability in Erodium. Ann Bot 117:1209–1220. https://doi.org/10.1093/aob/mcw065
Calvente A, Moraes EM, Lavor P, Bonatelli IAS, Nacaguma P, Versieux LM, Taylor NP, Zappi DC (2016) Phylogenetic analyses of Pilosocereus (Cactaceae) inferred from plastid and nuclear sequences. Bot J Linn Soc 183:25–38. https://doi.org/10.1111/boj.12491
Cauz-Santos LA, da Costa ZP, Callot C, Cauet S, Zucchi MI, Bergès H, van den Berg C, Vieira MLC (2020) A repertory of rearrangements and the loss of an inverted repeat region in Passiflora chloroplast genomes. Genome Biol Evol. https://doi.org/10.1093/gbe/evaa155
Choi I-S, Jansen R, Ruhlman T (2019) Lost and found: return of the inverted repeat in the legume clade defined by its absence. Genome Biol Evol 11:1321–1333. https://doi.org/10.1093/gbe/evz076
Coradin L, Siminski A, Reis A (2011) Espécies Nativas da Flora Brasileira de Valor Econômico Atual ou Potencial: Plantas para o Futuro - Região Sul. MMA, Brasília
Darling ACE, Mau B, Blattner FR, Perna NT (2004) Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome Res 14:1394–1403. https://doi.org/10.1101/gr.2289704
Dierckxsens N, Mardulyn P, Smits G (2016) NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res 45:2–9. https://doi.org/10.1093/nar/gkw955
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Edwards EJ, Nyffeler R, Donoghue MJ (2005) Basal cactus phylogeny: implications of Pereskia (Cactaceae) paraphyly for the transition to the cactus life form. Am J Bot 92:1177–1188. https://doi.org/10.3732/ajb.92.7.1177
Fantinati MR, Soffiatti P, Calvente A (2021) A new phylogenetic hypothesis for Cereinae (Cactaceae) points to a monophyletic subtribe. Syst Bot 46:689–699. https://doi.org/10.1600/036364421X16312068417020
Ferreira ME, Grattapaglia D (1998). Introdução ao uso de marcadores moleculares em análise genética. (3ed). EMBRAPA-CENARGEN, Brasília
Gantt JS, Baldauf SL, Calie PJ, Weeden NF, Palmer JD (1991) Transfer of rpl22 to the nucleus greatly preceded its loss from the chloroplast and involved the gain of an intron. EMBO J 10:3073–3078. https://doi.org/10.1002/j.1460-2075.1991.tb07859.x
Greiner S, Lehwark P, Bock R (2019) OrganellarGenomeDRAW (OGDRAW) version 131: expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res. https://doi.org/10.1093/nar/gkz238
Guo X, Liu J, Hao G, Zhang L, Mao K, Wang X, Zhang D, Ma T, Hu Q, Al-Shehbaz IA, Koch MA (2017) Plastome phylogeny and early diversification of Brassicaceae. BMC Genomics 18:176. https://doi.org/10.1186/s12864-017-3555-3
Hernández-Hernández T, Hernandez HM, De-Nova JA, Puente R, Eguiarte LE, Magallon S (2011) Phylogenetic relationships and evolution of growth form in Cactaceae (Caryophyllales, Eudicotyledoneae). Am J Bot 98:44–61. https://doi.org/10.3732/ajb.1000129
Hernández-Hernández T, Brown JW, Schlumpberger BO, Eguiarte LE, Magallón S (2014) Beyond aridification: multiple explanations for the elevated diversification of cacti in the New World Succulent Biome. New Phytol 202:1382–1397. https://doi.org/10.1111/nph.12752
Hershkovitz MA, Zimmer EA (1997) On the evolutionary origins of the Cacti. Taxon 46:217–232. https://doi.org/10.2307/1224092
Hunt D, Taylor N, International Cactaceae Systematics Group (2006) The new cactus lexicon. dh Books, Milborne Port
IUCN (2021) IUCN red list categories and criteria: version 3.1, 2nd ed. IUCN, Gland, Switzerland and Cambridge, UK
Jansen RK, Saski C, Lee S-B, Hansen AK, Daniell H (2010) Complete plastid genome sequences of three rosids (Castanea, Prunus, Theobroma): evidence for at least two independent transfers of rpl22 to the nucleus. Mol Biol Evol 28:835–847. https://doi.org/10.1093/molbev/msq261
Ji Y, Yang L, Chase MW, Liu C, Yang Z, Yang J, Yang J-B, Yi T-S (2019) Plastome phylogenomics, biogeography, and clade diversification of Paris (Melanthiaceae). BMC Plant Biol 19:543. https://doi.org/10.1186/s12870-019-2147-6
Jin D-M, Wicke S, Gan L, Yang J-B, Jin J-J, Yi T-S (2020) The loss of the inverted repeat in the putranjivoid clade of malpighiales. Front Plant Sci 11:942. https://doi.org/10.3389/fpls.2020.00942
Judd WS, Campbell CS, Kellogg EA, Stevens PF, Donoghue MJ (2015) Plant systematics: a phylogenetic approach, 4th ed. Oxford University Press, New York
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Kim HT, Kim JS, Moore MJ, Neubig KM, Williams NH, Whitten WM, Kim J-H (2015) Seven new complete plastome sequences reveal rampant independent loss of the ndh gene family across orchids and associated instability of the inverted repeat/small single-copy region boundaries. PLoS One 10:11-e0142215. https://doi.org/10.1371/journal.pone.0142215
Kode V, Mudd EA, Iamtham S, Day A (2005) The tobacco plastid accD gene is essential and is required for leaf development. Plant J 44:237–244. https://doi.org/10.1111/j.1365-313X.2005.02533.x
Köhler M, Reginato M, Souza-Chies TT, Majure LC (2020) Insights into chloroplast genome evolution across opuntioideae (Cactaceae) reveals robust yet sometimes conflicting phylogenetic topologies. Front Plant Sci 11:729. https://doi.org/10.3389/fpls.2020.00729
Lin C-S, Chen JJW, Huang Y-T et al (2015) The location and translocation of ndh genes of chloroplast origin in the Orchidaceae family. Sci Rep 5:9040. https://doi.org/10.1038/srep09040
Liu X, Yang H, Zhao J, Zhou B, Li T, Xiang B (2017) The complete chloroplast genome sequence of the folk medicinal and vegetable plant purslane (Portulaca oleracea L.). J. Hortic. Sci. Biotechnol 93:356–365. https://doi.org/10.1080/14620316.2017.1389308
Lowe T, Chan P (2016) tRNAscan-SE On-line: integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res 44(W1):W54-W57. https://doi.org/10.1093/nar/gkw413
Lüttge U (2004) Ecophysiology of crassulacean acid metabolism (CAM). Ann Bot 93:629–652. https://doi.org/10.1093/aob/mch087
Martín M, Sabater B (2010) Plastid ndh genes in plant evolution. Plant Physiol Biochem 48:636–645. https://doi.org/10.1016/j.plaphy.2010.04.009
Martínez-Alberola F, del Campo EM, Lázaro-Gimeno D, Mezquita-Claramonte S, Molins A, Mateu-Andrés I, Pedrola-Monfort J, Casano LM, Barreno E (2013) Balanced gene losses, duplications and intensive rearrangements led to an unusual regularly sized genome in Arbutus unedo chloroplasts. PLoS ONE 8:11-e79685. https://doi.org/10.1371/journal.pone.0079685
Mauseth JD (2006) Structure-function relationships in highly modified shoots of cactaceae. Ann Bot 98:901–926. https://doi.org/10.1093/aob/mcl133
Mower JP, Vickrey TL (2018) Structural diversity among plastid genomes of land plants. In: Chaw SMJR (ed) Advances in botanical research. Academic Press, Cambridge, pp 263–292
Neuhaus HE, Emes MJ (2000) Nonphotosynthetic metabolism in plastids. Annu Rev Plant Physiol Plant Mol 51:111–140. https://doi.org/10.1146/annurev.arplant.51.1.111
Nevill PG, Howell KA, Cross AT, Williams AV, Zhong X, Tonti-Filippini J, Boykin LM, Dixon KW, Small I (2019) Plastome-wide rearrangements and gene losses in carnivorous Droseraceae. Genome Biol Evol 11:472–485. https://doi.org/10.1093/gbe/evz005
Nyffeler R (2002) phylogenetic relationships in the cactus family (Cactaceae) based on evidence from trnK/matK and trnL-trnF sequences. Am J Bot 89:312–326. https://doi.org/10.3732/ajb.89.2.312
Nyffeler R (2007) The closest relatives of cacti: insights from phylogenetic analyses of chloroplast and mitochondrial sequences with special emphasis on relationships in the tribe Anacampseroteae. Am J Bot 94:89–101. https://doi.org/10.3732/ajb.94.1.89
Oulo MA, Yang J-X, Dong X et al (2020) Complete chloroplast genome of Rhipsalis baccifera, the only cactus with natural distribution in the old world: genome rearrangement, intron gain and loss, and implications for phylogenetic studies. Plants 9:979. https://doi.org/10.3390/plants9080979
Palmer JD, Thompson WF (1982) Chloroplast DNA rearrangements are more frequent when a large inverted repeat sequence is lost. Cell 29:537–550. https://doi.org/10.1016/0092-8674(82)90170-2
Peltier G, Cournac L (2002) Chlororespiration. Annu Rev Plant Biol 53:523–550
Pérez-Molphe-Balch E, MdelS S-D, Ramírez-Malagón R, Ochoa-Alejo N (2015) Tissue culture of ornamental cacti. Sci Agric 72:540–550. https://doi.org/10.1590/0103-9016-2015-0012
Ritz CM, Martins L, Mecklenburg R, Goremykin V, Hellwig FH (2007) The molecular phylogeny of Rebutia (Cactaceae) and its allies demonstrates the influence of paleogeography on the evolution of South American mountain Cacti. Ann Bot 94:1321–1322. https://doi.org/10.3732/ajb.94.8.1321
Rogalski M, Ruf S, Bock R (2006) Tobacco plastid ribosomal protein S18 is essential for cell survival. Nucleic Acids Res 34:4537–4545. https://doi.org/10.1093/nar/gkl634
Romani I, Tadini L, Rossi F, Masiero S, Pribil M, Jahns P, Kater M, Leister D, Pesaresi P (2012) Versatile roles of Arabidopsis plastid ribosomal proteins in plant growth and development. Plant J 72:922–934. https://doi.org/10.1111/tpj.12000
Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC, Guirao-Rico S, Librado P, Ramos-Onsins SE, Sánchez-Gracia A (2017) DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol Biol Evol 34:3299–3302. https://doi.org/10.1093/molbev/msx248
Ruhlman TA, Jansen RK (2018) Aberration or analogy? the atypical plastomes of geraniaceae. In: Chaw SMJR (ed) Advances in botanical research. Academic Press, Cambridge, pp 223–262
Sanderson MJ, Copetti D, Burquez A et al (2015) Exceptional reduction of the plastid genome of saguaro cactus (Carnegiea gigantea): loss of the ndh gene suite and inverted repeat. Am J Bot 102:1115–1127. https://doi.org/10.3732/ajb.1500184
Santana Neto JA, Castro Filho ES, Araújo HR (2015) Potencial das cactáceas como alternativa alimentar para ruminantes no semiárido. Revista Eletrônica Nutritime 12:4426–4434
Santiago RAC, Coradin L (2018) Biodiversidade Brasileira: sabores e aromas. MMA, Brasília
Santos DC, Farias I, Lira MA, Santos MVF, Arruda GP, Coelho RSB, Dias FM, Melo JN (2006) Manejo e utilização da palma forrageira (Opuntia e Nopalea) em Pernambuco. IPA, Recife
Silva GAR, Antonelli A, Lendel A, EdeM Moraes, Manfrin MH (2017) The impact of early Quaternary climate change on the diversification and population dynamics of a South American cactus species. J Biogeogr 45:76–88. https://doi.org/10.1111/jbi.13107
Silva GM, Lopes AS, Pacheco TG et al (2021) Genetic and evolutionary analyses of plastomes of the subfamily Cactoideae (Cactaceae) indicate relaxed protein biosynthesis and tRNA import from cytosol. Braz J Bot 44:97–116. https://doi.org/10.1007/s40415-020-00689-2
Solórzano S, Chincoya DA, Sanchez-Flores A, Estrada K, Díaz-Velásquez CE, González-Rodríguez A, Vaca-Paniagua F, Dávila P, Arias S (2019) De Novo assembly discovered novel structures in genome of plastids and revealed divergent inverted repeats in Mammillaria (Cactaceae, Caryophyllales). Plants 8:392. https://doi.org/10.3390/plants8100392
Souza DD, Pacheco CSGR (2019) Espécies nativas para alimentação de ruminantes em Ouricuri-PE e seus impactos ambientais. Agropecuária Científica no Semiárido 15:71–77. https://doi.org/10.30969/acsa.v15i1.1108
Taylor NP (1991) The genus Melocactus in Central and South America. Bradleya 9:1–80. https://doi.org/10.25223/brad.n9.1991.a1
Taylor N, Zappi D (2004) Cacti of Eastern Brazil. Royal Botanic Gardens, Kew, London
Taylor N, Zappi D (2018) Additions and corrections to “Cacti of Eastern Brazil.” Bradleya 36:2–21. https://doi.org/10.25223/brad.n36.2018.a2
Taylor N, Meiado MV, Filho EB, Zappi D (2014) A new Melocactus from the Brazilian state of Sergipe. Bradleya 32:99–104. https://doi.org/10.25223/brad.n32.2014.a3
Tesler G (2002) GRIMM: genome rearrangements web server. Bioinformatics 18:492–493. https://doi.org/10.1093/bioinformatics/18.3.492
Tillich M, Lehwark P, Pellizzer T, Ulbricht-Jones ES, Fischer A, Bock R, Greiner S (2017) GeSeq – versatile and accurate annotation of organelle genomes. Nucleic Acids Res 45:W6–W11. https://doi.org/10.1093/nar/gkx391
Ueda M, Kuniyoshi T, Yamamoto H, Sugimoto K, Ishizaki K, Kohchi T, Nishimura Y, Shikanai T (2012) Composition and physiological function of the chloroplast NADH dehydrogenase-like complex in Marchantia polymorpha. Plant J 72:683–693. https://doi.org/10.1111/j.1365-313x.2012.05115.x
Weng M-L, Ruhlman TA, Jansen RK (2016) Expansion of inverted repeat does not decrease substitution rates in Pelargonium plastid genomes. New Phytol 214:842–851. https://doi.org/10.1111/nph.14375
Wicke S, Naumann J (2018) Molecular evolution of plastid genomes in parasitic flowering plants. In: Chaw SMJR (ed) Advances in botanical research. Academic Press, Cambridge, pp 315–347
Wicke S, Schneeweiss GM, dePamphilis CW, Müller KF, Quandt D (2011) The evolution of the plastid chromosome in land plants: gene content, gene order, gene function. Plant Mol Biol 76:273–297. https://doi.org/10.1007/s11103-011-9762-4
Yao G, Jin J-J, Li H-T et al (2019) Plastid phylogenomic insights into the evolution of Caryophyllales. Mol Phylogenet Evol 134:74–86. https://doi.org/10.1016/j.ympev.2018.12.023
Acknowledgements
The authors thank the INSA (Instituto Nacional do Semiárido) for donation of the specimens analysed, André Marques (Max Planck Institute for Plant Breeding Research (MPIPZ), Köln, Germany) for assistance with the use of NOVOPlasty and Marlon Machado for suggestions at initial stages of the project.
Funding
The authors thank FACEPE (Fundacão de Amparo à Ciência e Tecnologia de Pernambuco) [IBPG-1030–2.03/17] (EM) and CNPq (Conselho Nacional de Desenvolvimento Científco e Tecnológico) (AP-H) for the grants awarded. This study was supported in part by the Coordenacão de Aperfeiçoamento de Pessoal de Nıvel Superior—Brasil (CAPES, Finance Code 001).
Author information
Authors and Affiliations
Contributions
EMA conducted the experiments, performed data analysis, and wrote the manuscript; MAS performed bioinformatic analysis, analysed data, and assisted with writing the manuscript; PER helped with DNA extraction, plant cultivation, and discussion of data; BL helped with bioinformatic analysis and assisted with writing the manuscript; LPF collected plant material and discussed data; APH designed the experiments and assisted with writing the manuscript. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
de Almeida, E.M., Sader, M.A., Rodriguez, P.E. et al. Assembling the puzzle: Complete chloroplast genome sequences of Discocactus bahiensis Britton & Rose and Melocactus ernestii Vaupel (Cactaceae) and their evolutionary significance. Braz. J. Bot 44, 877–888 (2021). https://doi.org/10.1007/s40415-021-00772-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40415-021-00772-2