Abstract
In most eukaryotic species, centromeres harbor large arrays of tandem repeated satellite DNA sequences. In this study, we report on the genomic distribution of a centromere satellite repeat “MtR3” in Medicago genus and three distantly related genera. Fluorescence in situ hybridization (FISH) results showed MtR3 repeats were detected in the centromere regions in M. truncatula, M. minima, M. edgeworthii, M. ruthenica, M. caerulea, M. sativa, and M. falcata (4×), but no signals were discovered in M. lupulina, M. polymorpha, and M. falcata (2×), Melilotus officinalis, Crotalaria medicaginea, and Trifolium repens. However, sequence analysis showed this MtR3 DNA had genomic distribution in all species and was highly conserved across the entire Medicago genus and three other genera. The conservation and widespread presence suggested MtR3 repeats may play important roles in centromeric function.





Similar content being viewed by others
References
Allen G, Flores-Vergara M, Krasynanski S, Kumar S, Thompson W (2006) A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nat Protoc 1:2320–2325
Alves G, Seuánez HN, Fanning T (1994) Alpha satellite DNA in neotropical primates (Platyrrhini). Chromosoma 103:262–267
Ananiev E, Phillips R, Rines H (1998) Chromosome-specific molecular organization of maize (Zea mays L.) centromeric regions. Proc Natl Acad Sci USA 95:13073–13078
Ansari HA, Ellison NW, Griffiths AG, Williams WM (2004) A lineage-specific centromeric satellite sequence in the genus Trifolium. Chromosome Res 12:357–367
Bauchan GR, Hossain MA (1997) Identification of Alfalfa chromosomes using Geimsa banding and image analysis techniques. In: Proceedings of the XVIII international rangeland congress, Canada, pp 61–62
Bena G (2001) Molecular phylogeny supports the morphologically based taxonomic transfer of the “medicagoid” Trigonella species to the genus Medicago L. Plant Syst Evol 229:217–236
Bena G, Lejeune B, Prosperi JM, Olivieri I (1998a) Molecular phylogenetic approach for studying life-history evolution: the ambiguous example of the genus Medicago L. Proc R Soc London, Ser B 265:1141–1151
Bena G, Jubier MF, Olivieri I, Lejeune B (1998b) Ribosomal external and internal transcribed spacers: combined use in the phylogenetic analysis of Medicago (Leguminosae). J Mol Evol 46:299–306
Calderini O, Pupilli F, Paolocci F, Arcioni S (1997) A repetitive and species-specific sequence as a tool for detecting the genome contribution in somatic hybrids of the genus Medicago. Theor Appl Genet 95:734–740
Charlesworth B, Sniegowski P, Stephan W (1994) The evolutionary dynamics of repetitive DNA in eukaryotes. Nature 371:215–220
Cheng Z et al (2002) Functional rice centromeres are marked by a satellite repeat and a centromere-specific retrotransposon. Plant Cell 14:1691–1704
Dou QW, Chen ZG, Liu YA, Tsujimoto H (2009) High frequency of karyotype variation revealed by sequential FISH and GISH in plateau perennial grass forage Elymus nutans. Breed Sci 59:651–656
Ferreira D, Meles S, Escudeiro A, Mendes-da-Silva A, Adega F, Chaves R (2015) Satellite non-coding RNAs: the emerging players in cells, cellular pathways and cancer. Chromosome Res 23:479–493
Havananda T, Brummer EC, Maureira-Butler IJ, Doyle JJ (2010) Relationships among diploid members of the Medicago sativa (Fabaceae) species complex based on chloroplast and mitochondrial DNA sequences. Syst Bot 35:140–150
Henikoff S, Ahmad K, Malik HS (2001) The centromere paradox: stable inheritance with rapidly evolving DNA. Science 293:1098–1102
Heslop-Harrison JS, Murata M, Ogura Y, Schwarzacher T, Motoyoshi F (1999) Polymorphisms and genomic organization of repetitive DNA from centromeric regions of Arabidopsis chromosomes. Plant Cell 11:31–42
Iwata A et al (2013) Identification and characterization of functional centromeres of the common bean. Plant J 76:47–60
Jiang J et al (1996) A conserved repetitive DNA element located in the centromeres of cereal chromosomes. Proc Natl Acad Sci USA 93:14210–14213
Kulikova O et al (2004) Satellite repeats in the functional centromere and pericentromeric heterochromatin of Medicago truncatula. Chromosoma 113:276–283
Lamb JC, Yu W, Han F, Birchler JA (2007) Plant chromosomes from end to end: telomeres, heterochromatin and centromeres. Curr Opin Plant Biol 10:116–122
Ma J, Wing RA, Bennetzen JL, Jackson SA (2007) Plant centromere organization: a dynamic structure with conserved functions. Trends Genet 23:134–139
Maureira-Butler IJ, Pfeil BE, Muangprom A, Osborn TC, Doyle JJ (2008) The reticulate history of Medicago (Fabaceae). Syst Biol 57:466–482
Mehrotra S, Goyal V (2014) Repetitive sequences in plant nuclear DNA: types, distribution, evolution and function. Genom Proteom Bioinform 12:164–171
Melters DP et al (2013) Comparative analysis of tandem repeats from hundreds of species reveals unique insights into centromere evolution. Genome Biol 14:R10
Mestrović N, Plohl M, Mravinac B, Ugarković D (1998) Evolution of satellite DNAs from the genus Palorus-experimental evidence for the “library” hypothesis. Mol Biol Evol 15:1062–1068
Plohl M, Luchetti A, Meštrović N, Mantovani B (2008) Satellite DNAs between selfishness and functionality: structure, genomics and evolution of tandem repeats in centromeric (hetero) chromatin. Gene 409:72–82
Plohl M, Meštrović N, Mravinac B (2014) Centromere identity from the DNA point of view. Chromosoma 123:313–325
Quiros CF, Bauchan GR (1988) The genus Medicago and the origin of the Medicago sativa complex. In: Hanson AA, Barnes DK, Hill RR (eds). Alfalfa and alfalfa improvement. American Society of Agronomy, Crop Science Society of America, Soil Science Society of America, Madison, pp 93–124
Rosato M, Galián JA, Rosselló JA (2012) Amplification, contraction and genomic spread of a satellite DNA family (E180) in Medicago (Fabaceae) and allied genera. Ann Bot 109:773–782
Rošić S, Köhler F, Erhardt S (2014) Repetitive centromeric satellite RNA is essential for kinetochore formation and cell division. J Cell Biol 207:335–349
Small E, Jomphe M (1989) A synopsis of the genus Medicago (Leguminosae). Can J Bot 67:3260–3294
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Topp CN, Zhong CX, Dawe RK (2004) Centromere-encoded RNAs are integral components of the maize kinetochore. Proc Natl Acad Sci USA 101:15986–15991
Ugarković Ð, Plohl M (2002) Variation in satellite DNA profiles-causes and effects. EMBO J 21:5955–5959
Wang G, Zhang X, Jin W (2009) An overview of plant centromeres. J Genet Genom 36:529–537
Willard HF (1990) Centromeres of mammalian chromosomes. Trends Genet 6:410–416
Yu F, Lei Y, Li Y, Dou Q, Wang H, Chen Z (2013) Cloning and characterization of chromosomal markers in alfalfa (Medicago sativa L.). Theor Appl Genet 126:1885–1896
Acknowledgements
We thank Professor Tao Wang and Jiangli Dong (State Key Laboratory of Agrobiotechnology, College of Biological Sciences, China Agricultural University, Beijing 100193, China) for providing partial Medicago materials. This work was supported by the Science and Technology Service Network Initiative of the Chinese Academy of Sciences (KFJ-SW-STS-177) and Natural Science Foundation of Qinghai Province (No. 2015-ZJ-903).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Feng Yu, Quanwen Dou, Ruijuan Liu, Haiqing Wang declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human subjects or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
13258_2017_556_MOESM2_ESM.tif
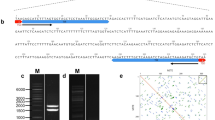
Suppl. Fig. S2 DNA sequence alignment of all MtR3 repeat units from 10 Medicago species and three related species. Blue shadow boxes indicate sequence identity above 50% homology level. Pink shadow boxes indicate sequence identity above 75% homology level. Black shadow boxes indicate sequence identity at 100% homology level. Consensus sequence is presented in the last row (TIF 2369 KB)
Rights and permissions
About this article
Cite this article
Yu, F., Dou, Q., Liu, R. et al. A conserved repetitive DNA element located in the centromeres of chromosomes in Medicago genus. Genes Genom 39, 903–911 (2017). https://doi.org/10.1007/s13258-017-0556-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-017-0556-1