Abstract
The Dalmatian Daisy Tanacetum cinerariifolium is an Asteraceae plant species that produces the natural insecticide “pyrethrum”, which is effective against mosquito disease vectors and household pests. To enhance the content of pyrethrum in flowers, a more detailed understanding of the mechanisms underlying pyrethrum biosynthesis is needed. Even though gene transformation and genome editing techniques are vital for investigating pyrethrin biosynthesis, limited information is available on the transformation of T. cinerariifolium. Furthermore, each seedling possesses a distinct genotype with large variations by self-incompatibility. We herein employed T. cinerariifolium line #14 with weak self-incompatibility to establish a protocol of efficient regeneration from leaf segments and transformation. Leaf segments formed calli on 1/2 Murashige and Skoog’s basal medium (MS) with naphthalene acetic acid 1 mg L−1 and 6-benzylaminopurine (BAP) 2 mg L−1, regenerated shoots from calli on 1/2 MS with BAP 0.5 mg L−1 and GA3 0.2 mg L−1, and elongated shoot stems on 1/2 MS with indole-3-butyric acid 0.5 mg L−1 and BAP 0.5 mg L−1. To establish genetic transformation, Rhizobium radiobacter strain EHA105 with the highest infectivity and the mas1'-2' bidirectional promoter with the highest expression of the nptII resistance gene were used, and the antibiotic G418 was added to medium at a concentration of 10 to 20 mg L−1 to select transformed cells. Using established regeneration techniques, we successfully obtained transformants that highly expressed the transgene gusA. This technique will be useful for creating genetically modified T. cinerariifolium, particularly for elucidating the mechanism of pyrethrin biosynthesis toward the creation of pyrethrin-rich traits.
Key messages
A reproducible genetic transformation system is not yet available for Tanacetum cinerariifolium. We created a transformation technique using a Rhizobium radiobacter strain and a vector for high gene expression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Dalmatian Daisy Tanacetum cinerariifolium is an Asteraceae plant species that produces pyrethrins, which are natural insecticides. Its dried flower extract, “Pyrethrum”, exhibits high knockdown activity against various pest insect species by modulating neural transmission (Matsuda et al. 2005; Lybrand et al. 2020; Varga et al. 2022). Pyrethrum is effective against mosquito disease vectors and household pests. The active components pyrethrins modulate voltage-sensitive sodium channels, thereby inducing repetitive firing and the subsequent blockade of neural conduction (Narahashi 1992). Recent studies reported that pyrethrins interacted with odorant receptors in antennae and induced repellency in flying insects (Liu et al. 2021; Wang et al. 2021). While the sale of pyrethrins is increasing, supply is limited, as is the case with natural products of plant origin that are subject to changes in temperature and rain fall as well as disease infection and herbivory. A more detailed understanding of the mechanisms underlying pyrethrin biosynthesis is needed to increase pyrethrin contents in flowers.
The genome of Tanacetum cinerariifolium is principally diploid but may become triploid and tetraploid by repeated crossing. The genome size is ~ 7 Gb and its sequence has been partially elucidated (Yamashiro et al. 2019). Several biosynthetic enzymes have been identified. However, the roles of most gene products and their regulation remain largely unknown. Self-incompatibility and the difficulties associated with genetically modifying this species are obstacles to elucidating the mechanisms underlying pyrethrin biosynthesis.
Previous studies reported methods for tissue cultivation (Roest and Bokeimann 1973; Aoki et al. 1976; Cashyap et al. 1978; Hitmi et al. 1999; Hedayat et al. 2009) and genetic transformation (Li et al. 2022a, b; Mao et al. 2013). However, each study used different culture methods and transgenic techniques; these methods have not been used in the development of new transgenic lines of T. cinerariifolium, which may be attributed to its self-incompatibility and each seedling possessing a distinct genotype with large variations (Bhat and Menary 1986; Parlevliet 1974). Furthermore, shoot regeneration competence varies among genotypes in tissue cultures (Teixeira da Silva 2003). Therefore, the development of an easy, reproducible and efficient genetic transformation technique is needed. Here we report a protocol for efficiently transforming T. cinerariifolium material #14 via Rhizobium radiobacter (Agrobacterium tumefaciens).
Materials and Methods
Plant materials and preculture conditions
Pyrethrum (Tanacetum cinenariifolium) #14 with weak self-compatibility, found at Innoshima Island in Hiroshima, Japan, was used in experiments. Seeds were surface-sterilized by briefly dipping in 70% ethanol for thirty seconds and then in a 1% sodium hypochlorite solution (including 1% Tween 20) for 15 min, followed by rinsing 3 times for 10 min in sterile distilled water. Sterilized seeds were sown in plastic Petri dishes (10 cm in diameter) on Murashige and Skoog’s (1962) basal medium (Murashige and Skoog 1962) (MS; containing 3% sucrose). One week later, germinated seeds were replanted in plant culture boxes (Bio Medical Science, Co., Japan) containing four different media: MS (containing 3% sucrose), 1/2 MS medium containing half-strength inorganic nutrients (1/2 MS; containing 1.5% sucrose), White medium containing 2% sucrose (White 1963), and Gamborg’s B5 medium containing 2% sucrose (Gamborg et al. 1968). Each medium was adjusted to pH 5.8 and 0.3% Gellan Gum (Gel.) (Wako Pure Chemical Industries, Ltd., Japan) or 0.8% Agar (Wako Pure Chemical Industries, Ltd., Japan) was added prior to autoclaving at 120 °C for 15 min. Cultures were kept at 25 °C under a 16-h photoperiod using cool white fluorescent lamps or at 25 °C in darkness. The lamps provided a photosynthetic photon flux [PPF (400-700 nm)] of 60 μmol m−2 s−1. After two months, rooting frequency, plant height, and root length were measured. Ten germinated seeds were replanted in plant culture boxes containing the respective media and data were collected on 10 replicates.
Callus induction and regeneration
The effects of the material leaf growth stage and the combinations of phytohormones on regeneration efficiency were examined. Leaves in the following growth stages were used in this experiment: Age#1, leaves just expanding (leaf length of 3 to 5 mm); Age#2, leaves in the first week after expanding (leaf length of approximately 8 mm to 1 cm); and Age#3, leaves in the second week after expanding (leaf length of 1 to 1.5 cm) (See Fig. 1). The phytohormone 6-benzylaminopurine (BAP) as a cytokinin and 1-naphthylacetic acid (NAA) as an auxin were used to regulate regeneration from leaf segments. Leaf segments were inoculated on 1/2 MS solid medium supplemented with 9 combinations of BAP (0.5, 1.0, and 2.0 mg L−1) and NAA (0.1, 0.5, and 1.0 mg L−1). After three weeks, the number of leaf segments that formed calli or shoots was counted. The green callus (length < 1 mm) was inoculated on seven different regeneration medium [1/2 MS solid medium + 2.0 mg L−1 BAP + 1.0 mg L−1 NAA; 1/2 MS solid medium + 0.5, 1.0, and 2.0 mg L−1 BAP; and 0.1 and 0.2 mg L−1 gibberellic acid (GA3)] and each shoot length was measured three weeks after replacement. Twelve leaf fragments or green calli were placed in Petri dishes containing the respective media and data were collected on 10 replicates.
Leaf ages and cutting sites for leaf segments of Tanacetum cinerariifolium. Age#1: leaves just expanding, with a leaf length of 3 to 5 mm; Age#2: leaves in the first week after expanding, with a leaf length of approximately 8 mm to 1 cm; Age#3: leaves in the second week after expanding, with a leaf length of 1 to 1.5 cm. The scale bar indicates 10 mm. Red bars indicate cutting sites for leaf segments
Elongated shoots (length > 3 mm) were placed in four different stem elongation media: 1/2 MS solid medium + BAP (0.0 and 0.5 mg L−1) and indole-3-butyric acid (IBA; 0.5 and 1.0 mg L−1). Shoots that formed elongated stems were placed in phytohormone-free 1/2 MS solid medium for rooting. Each medium contained 1.5% sucrose, except for stem elongation and rooting media, which contained 1%. pH was adjusted to 5.8 and 0.3% Gel. was added prior to autoclaving at 120 °C for 15 min. Cultures were placed at 25 °C under a 16-h photoperiod using cool white fluorescent lamps or at 25 °C in darkness. The lamps provided PPF (400–700 nm) of 60 μmol m−2 s−1. Four shoots were replanted in plant culture boxes containing the respective media and data were collected on 10 replicates.
Effects of Rhizobium strains and promoters on efficient gene insertion, selection, and stable expression of foreign genes
R. radiobacter viz. EHA105 (provided by Dr. L. S. Melchers, Zeneca Mogen) and LBA4404 (Ooms et al. 1982) were used in experiments. Four different binary vectors were used: pBIK201G (provided by Dr. H. Ichikawa, Institute of Agrobiological Sciences, National Agriculture and Food Research Organization: NARO), pBI121 (Jefferson 1987), and pBE2133 and pBE7133 (provided by Dr. I. Mitsuhara, NARO) (Mitsuhara et al. 1996) (see Fig. 2). In each vector, the gusA gene was controlled by a different promoter. In binary vector pBIK201G, the gusA gene and selectable marker gene nptII were driven by the novel bi-directional promoter fragment (465 bp) for the mannopine synthase-1' and -2' (mas1'-2') genes from R. radiobacter strain AtC1 (MAFF301276) provided by the NIAS Genebank (Tsukuba, Japan). In binary vector pBI121, the gusA gene was driven by the promoter region of the Cauliflower mosaic virus 35S RNA (CaMV 35S) promoter, which has been widely used as a constitutive strong promoter in plant transformation. In binary vector pBE2113 (Mitsuhara et al. 1996), the gusA gene was driven by the El2Ω promoter with two tandem repeats of the 5’-upstream sequence of the CaMV 35S promoter (-419 to -90), the G-free sequence (Ω sequence) in the 57-untranslated region of Tobacco mosaic virus. In binary vector pBE7133 (Mitsuhara et al. 1996), the gusA gene was driven by the E7Ω promoter with seven tandemly repeated enhancer-like elements (-90 to -290, relative to the site of the initiation of transcription as + 1) of the CaMV 35S promoter (E7) and the Ω sequence. In pBE2113 and pBE7133, GUS activity levels were 20- to 70-fold higher than those with the 35S promoter in pBI121 in Tobacco (Nicotiana tabacum cv. Samsun NN) and rice (Oryza sativa cv. Nipponbare) protoplasts (Mitsuhara et al. 1996). In pBI121, pBE2133-GUS, and pBE7133-GUS, the selectable marker gene nptII was driven by the nopaline synthase (nos) promoter from Rhizobium Ti plasmids (Bevan et al. 1983).
Structures of tested binary vectors. T35S, the polyadenylation signal of the Cauliflower mosaic virus (CaMV) 35S transcript; nptII, the neomycin phosphotransferase II gene for the selection of transgenic plants; Pmas201, the bidirectional promoter cassette from the mannopine synthase 1′ and 2′ (mas1′-2′) genes; gusA, the β-glucuronidase gene; Tnos, the polyadenylation signal of the gene for nopaline synthase in the Ti plasmid; Pnos, the nopaline synthase promoter from Rhizobium Ti plasmids; P35S, the 5’-upstream sequence of the CaMV 35S promoter (-90 to -1); E7, the 5’-upstream sequence of the CaMV 35S promoter (-940 to -290) and (-290 to -90) × 7; Ω, the 5’-upstream sequence of Tobacco mosaic virus (TMV); In, the 1st intron of the phageolin gene; El2, the 5’-upstream sequence of the CaMV 35S promoter (-419 to -90) × 2. Red arrows indicate gusA-specific primers to amplify an approximately 1.8-kbp fragment in the gusA coding region. Green arrows indicate nptII-specific primers to amplify an approximately 800-bp fragment in the nptII coding region. The blue bar indicates the nptII-specific probe for the Southern blot analysis
Each binary vector was transformed into Rhizobium strains EHA105 and LBA4404 using the triparental mating method (Ditta et al. 1980).
Rhizobium infection
The genetic transformation procedure reported by Shinoyama et al. (2002) was used with minor modifications (Table 1). R. radiobacter was cultured for 4 to 5 h in YEP liquid medium on Bio-Shaker BR-3000L (TAITEC, Japan) at 28 °C under dark conditions. Leaf segments were prepared by cutting with a scalpel, as shown in Fig. 1. They were immersed at room temperature for 15 min in Infection medium [1/2 MS liquid medium containing 5% Tween 20 (Nacalai Tesque, Japan) and 50 μM acetosyringone] with Rhizobium (final OD660 = 0.1). After immersion, leaf segments were placed onto co-cultivation medium [1/2MS solid medium + 1.0 mg L−1 NAA and 2.0 mg L−1 BAP (Callus formation solid medium; CS) containing 1.0 g L−1 casamino acid] and co-cultivated at 25 °C in darkness for 2 days.
Optimization of cefotaxime and G418 concentrations for Rhizobium elimination and transformant selection
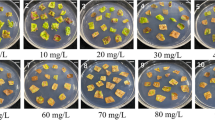
Two days after infection, leaf fragments were transferred to Bacteria elimination liquid medium (Table 1) supplemented with each concentration of cefotaxime sodium salt (250 and 500 mg L−1) to 1/2 MS liquid medium containing 1.0 mg L−1 NAA and 2.0 mg L−1 BAP (Callus formation liquid medium; CL) with shaking at 25 °C overnight under a 16-h photoperiod (Table 1). They were then placed in Bacteria elimination solid medium (CS + 100 mg L−1 cefotaxime sodium salt). Fourteen days after infection, leaf segments were transferred to Bacteria elimination liquid medium (CL + 100 mg L−1 cefotaxime sodium salt) with shaking at 25 °C overnight under a 16-h photoperiod. They were then placed in Selection medium (CS + 100 mg L−1 cefotaxime sodium salt and 10 or 20 mg L−1 G418) for the selection of putative transformed callus. Leaf fragments were transferred to new medium every 14 days. The number of green calli was counted after transplanting to the selection medium twice at an interval of 14 days.
Analysis of gusA gene expression
Leaf explants, calli, regenerated shoots, and regenerated plants were confirmed by the GUS histochemical assay according to Jefferson (1987) and Murakami and Ohashi (1992). Calli and regenerated shoots were incubated in 50 mM phosphate buffer (pH 7.2) containing 1 mM 5-bromo-4-chloro-3-indoyl glucuronide (X-Gluc), 5 mM dithiothreitol (DTT), 0.3% Triton X-100, 5% methanol, 0.5 mM potassium ferrocyanide, and 0.5 mM potassium ferricyanide at 37 °C overnight. After staining, calli and regenerated shoots were decolorized with 99% ethanol for the extraction of chlorophyll. GUS activity in transgenic callus was analyzed by the fluorometric method reported by Kosugi et al. (1990) with 4-methyl-umbelliferyl glucuronide as the substrate. Twelve leaf segments were used in the GUS assay (number of blue spots) and 10 replications were performed. In the GUS activity test, ten calli and shoots were used, and data were collected on 10 replications.
DNA extraction and polymerase chain reaction (PCR) analysis of the nptII gene and gusA gene
Total DNA was extracted from 100 mg of the fresh young leaves of regenerated or control plantlets by the method of Takagi et al. (1993). Leaves were homogenized in liquid nitrogen using a ceramic mortar and pestle and suspended in 1 ml of HEPES buffer [0.1 M HEPES (pH 8.0), 0.1% PVP, and 4% 2-mercaptoethanol]. After centrifugation at 15,000 rpm at 4 °C for 5 min, the supernatant was discarded, and the pellet was suspended in a new HEPES buffer. This procedure was repeated 3 times to remove polyphenols and polysaccharides. Total DNA was isolated from the pellet by the sodium dodecyl sulfate extraction method reported by Honda and Hirai (1990). PCR was performed using 200 ng genomic DNA, 1 U recombinant Taq DNA Polymerase (TaKaRa Taq™, Takara Bio, Japan), 0.5 mM of specific forward and reverse primers, 0.2 mM dNTPs, and PCR buffer in a final volume of 20 μL. A PCR analysis was performed using the forward primer 5’-ATGTTACGTCCTGTAGAAAC-3’ and reverse primer 5’-TTCATTGTTTGCCTCCCTGC-3’ for the gusA gene (1,811 bp), and the forward primer 5’-ATGATTGAACAAGATGGATT-3’ and reverse primer 5’-TCAGAAGAACTCGTCAAGAA -3’ for the nptII gene (795 bp). Amplification was performed in Mastercycler® ep (Eppendorf SE, Japan) under the following conditions: pre-denaturation at 94 °C for 2 min, followed by 30 cycles (94 °C, 30 s; 55 °C, 30 s; 72 °C, 1 min), and finally at 72 °C for 5 min followed by an incubation at 4 °C. PCR products were separated on 1.0% (w/v) agarose gel, stained with ethidium bromide, and visualized under UV for documentation.
Southern blot hybridization
Total DNA was extracted from 100 mg of fresh young leaves of the transformed lines and non-transformed control plants according to Shinoyama et al. (2002). A 25-μg aliquot of DNA digested with XbaI was subjected to electrophoresis and blotted onto a Hybond N + nylon membrane (Sativa, USA). Southern blot hybridization (Southern 1975) was performed using a digoxigenin (DIG)-labeled nptII fragment (approximately 800 bp) as a probe (Fig. 2) and a DIG DNA Labeling and Detection Kit (Roche Diagnostics, Germany) according to the supplier’s instructions.
Statistical analysis
Data obtained from all experiments were presented as the mean ± SD and separation was performed using Turkey-Kramer’s HDS. Percentage data were transformed to arcsine data and statistical analyses were performed.
Results
Plant materials and preculture conditions
Ten germinated seeds of T. cinerariifolium were replanted in 4 kinds of media, which differed in solidifying agents and MS medium concentrations, and the growth of seedlings was examined. The rooting frequency ranged between 28.0 ± 0.4 and 80.0 ± 0.0% and was the highest in 1/2 MS + Gel. medium. Plant heights ranged between 14.7 ± 0.2 and 27.4 ± 1.0 cm, and were significantly higher on MS + Gel., MS + Agar, and 1/2 MS + Gel. medium than on other media. Root lengths ranged between 31.7 ± 0.7 and 52.4 ± 3.4 mm, with the highest being measured on Gamborg’s B5 + Gel. (Table 2).
Green callus induction and regeneration
Age #2 leaves were initially used as plant material for phytohormone combination tests. Green calli formed on the edge of segments after 3 weeks and some adventitious shoots formed on calli after 4 to 5 weeks. The number of green callus/shoot-forming leaf segments was measured and the green callus/shoot-forming leaf segment ratio per number of leaf segments tested was calculated (Table 3). The green callus formation ratio was significantly higher (Tukey–Kramer test, p < 0.01) on media containing 1.0 mg L−1 NAA + 0.5, 1.0, and 2.0 mg L−1 BAP and that containing 0.5 mg L−1 NAA + 1.0 mg L−1 BAP. Shoots formed on media containing 0.5 mg L−1 NAA + 1.0 mg L−1 BAP and 1.0 mg L−1 NAA + 2.0 mg L−1 BAP, with no significant difference in the formation ratio of both. Differences in the age of the leaves used as material (Age#1, #2, and #3, see Fig. 1) were examined using the medium with the highest callus formation ratio, namely, 1/2 MS + 1.0 mg L−1 NAA and 2.0 mg L−1 BAP medium. Formation ratios were significantly higher for Age#1 and #2 than for Age#3. No significant difference was observed between Age#1 and #2 (Table 4).
Green calli (length < 1 mm) were transferred to regeneration medium for shoot regeneration. The longest elongated length of 5.8 ± 4.0 mm was observed at 0.5 mg L−1 BAP + 0.2 mg L−1 GA3 (Table 5). In the combination of BAP and GA3, the elongation length slightly decreased at higher concentrations of BAP and increased at higher concentrations of GA3 (see Fig. 3). Shoots of approximately 3 mm were replanted in plant culture boxes with air vent membranes (Milliseal Membrane Seal, FWMS01800, Merck KGaA, Germany) and containing four different stem-elongating media (1/2 MS solid medium + BAP 0.0 and 0.5 mg L−1, IBA 0.5 and 1.0 mg L−1, respectively). Two weeks after plating, all shoots on 1/2 MS medium containing only 1.0 mg L−1 of IBA turned brown, whereas those placed on other media grew and formed stems (Fig. 4). Four weeks after plating, the medium with 1.0 mg L−1 IBA + 0.5 mg L−1 BAP resulted in the highest green callus formation ratio (Table 6). Shoots with elongated stems placed in phytohormone-free 1/2 MS medium showed rooting (Fig. 5).
Rooting from elongated shoots. Shoots with elongated stems on 1/2 MS medium containing 1.0 mg L−1 IBA and 0.5 mg L−1 BAP were replanted in 1/2 MS phytohormone-free medium. a: Shoots on 1/2 MS medium containing 1.0 mg L−1 IBA and 0.5 mg L−1; BAP; b: replanting shoots with elongated stems in 1/2 MS phytohormone-free medium; c: rooting shoots three or four weeks after replanting. Scale bars indicate 2 cm
Effects of Rhizobium strains and promoters for efficient insertion into the T. cineraria genome, selection, and stable expression of foreign genes
To evaluate Rhizobium infection ability and promoter activity, a histochemical assay for the gusA gene was performed three days after infection. Among the Rhizobium lines tested, the number of GUS spots was significantly higher at the 1% level for EHA105 than for LBA4404 for all promoters (Fig. 6a). GUS activity levels were analyzed using ten transgenic calli induced on selection medium containing 10 mg L−1 G418. On both calli and shoots, GUS activity was the highest at the mas1'-2' promoter carried by pBIK201, followed by the El2Ω promoter carried by pBE2113 and the E7Ω promoter carried by pBE7133, and was the lowest at the 35S promoter carried by pBI121 (Fig. 6b and 6c). GUS activity at the mas1'-2' promoter in pBIK201G was approximately sixfold higher than that at the 35S promoter in pBI121. No significant difference was observed in GUS activity between the El2Ω promoter and E7Ω promoter. Furthermore, GUS activity did not significantly differ among different Rhizobium strains with the same promoter. According to the histochemical GUS assay, chimeric expression was not detected in callus or adventitious shoots (Fig. 6d).
Effects of Rhizobium strains and promoters for the gusA gene on insertion and expression. (a): Effects of Rhizobium strains on efficient gene insertion. The GUS spot numbers of 12 leaf explants were measured with 10 replications. Each value represents the mean of GUS spots per leaf explant ± SD. (b): Effects of promoters on the efficient expression of foreign genes. The GUS activity of calli was measured with 10 replications. Each value represents mean GUS activity ± SD. (c): Effects of promoters on the efficient expression of foreign genes. The GUS activity of shoots was measured with 10 replications. Each value represents mean GUS activity ± SD. Values in a, b and c with the same letter were not significantly different at p < 0.01. (d): Effects of promoters on gusA gene expression in calli and regenerated shoots. The upper and lower figures represent expression in calli and regenerated shoots, respectively. Scale bars indicate 5 mm
Optimization of G418 concentrations for the efficient selection of transgenic lines
The elimination of Rhizobium performed 2 days after infection was possible at 250 or 500 mg L−1 of cefotaxime sodium salt in Bacteria elimination medium A and Rhizobium did not subsequently appear on the medium. Leaf segments 14 days after Rhizobium infection were replanted in selection medium containing 10 or 20 mg L−1 G418, and the number of calli was counted after transplanting to the selection medium twice at an interval of 14 days. The regeneration ratio was slightly higher on 500 mg L−1 than on 250 mg L−1 of cefotaxime sodium salt and higher on 10 mg L−1 than on 20 mg L−1 of G418. The highest regeneration ratio was 9.5% in the medium with 500 mg L−1 of cefotaxime sodium salt and 10 mg L−1 of G418 (Table 7). No band was detected in control plants by the PCR analysis. An approximately 800-bp single band of nptII and 1,800-bp single band of gusA were detected for some regenerated plants, but not for others (Supplementary Figure S1). Selection with 20 mg L−1 of G418 resulted in fewer regenerated plants than that with 10 mg L−1; notably, all the regenerated plants had transgenes (Table 7). The presence and number of T-DNA copies in individual regenerated plants were confirmed by the Southern blot analysis of XbaI-digested genomic DNA probed with a fragment of nptII. No hybridization signals were detected in non-transgenic T. cinerariifolium controls. Single to multiple bands hybridizing to the nptII probe were observed in regenerated plantlets, while no hybridization signals were detected in regenerated plants (Fig. 7a), which was consistent with the results of the PCR analysis (Supplementary Figure S1). The GUS assay on young leaves not fully expanded and the elongated roots of individuals confirmed the expression of the gusA gene (Fig. 7 c1 and c2) only in the transformants (Fig. 7b1 and b2).
Integration of the gusA gene into genomes of transgenic plants. (a): Southern blot analysis to detect transgene integration into the genomes of regenerated plants. A 25-μg aliquot of DNA was digested with XbaI and hybridized with the nptII-specific probe. M: λ-HindIII digest, P: the binary vector pBIK201G, N: non-transformed control, 1 to 7: regenerated plants with specific bands of gusA and nptII detected by PCR. b1 to c2: gusA gene expression in regenerated plants; b1 and c1: young leaves not fully expanded; b2 and c2: roots. A part of the photo enclosed by a rectangle is expanded to show a broad expression of the introduced gusA gene. Scale bars indicate 5 mm
Discussion
The callus and adventitious shoot formation technique is very useful for clonal propagation and the production of genetically modified plants. In T. cinerariifolium, Hitmi et al. (1999) induced adventitious shoots from calli, while Hedayat et al. (2009) and Mao et al. (2013) achieved regeneration via direct organogenesis from leaves. In the present study, we used T. cinerariifolium line #14 with weak self-compatibility for the callus formation, foreign gene introduction and regeneration of shoots. Difficulties were associated with the regeneration of this line from tissue due to browning (particularly leaf segments) during the tissue culture and a poor rooting ability from the shoots. Edible plants belonging to the Asteraceae family, such as burdock (Arctium lappa L.), Japanese butterbur (Petasites japonicus), Japanese mugwort (Artemisia indica var. maximowiczii), and dandelion (Taraxacum genus), undergo significant browning during cooking and processing (Han 2008). Regarding browning components, Nakamura (1968) identified polyphenols as chlorogens in burdock root, butterbur, and dandelion. Panizzi and Scarpati (1954) isolated an isomer of chlorogenic acid, cynarin (1.4-dicaffeyl quinic acid), from Cynarascolymus. Han (2008) demonstrated that Asteraceae vegetables contained chlorogenic acids as the main polyphenols, but at various contents, and that the enzymatic browning of Asteraceae vegetables resulted from the oxidative reaction of chlorogenic acids. Hence, we shook the culture and successfully reduced the browning of leaf segments, eliminated bacteria and performed a fast selection.
Hedayat et al. (2009) elongated shoots using BA-free B5 medium containing NAA only; however, this method did not work for shoot elongation from the #14 line. We found that the #14 line was rooted by the formation of fully elongated stems with IBA and BAP, which were then transferred to phytohormone-free 1/2 MS medium (Table 6 and Fig. 5). This rooting ability may be influenced by genetic backgrounds because of their self-incompatibility (Bhat and Menary 1986; Parlevliet 1974; Teixeira da Silva 2003).
For gene transformation efficiency, the number of GUS spots per leaf segment was threefold higher in EHA105 than in LBA4404. EHA105 is derived from super virulent strain A 281 (Hood et al. 1993), while LBA4404 is derived from the less virulent strain Ach 5 (Hoekema et al. 1983). EHA105 was previously reported to be more efficient than other strains for the transformation of Punica granatum (Terakami et al. 2007), Saccharum spp. (Manickavasagam et al. 2004), and Zea mays L. (Yu et al. 2013). On the other hand, the elimination of strain LBA4404 from plant tissues was easier using lower concentrations of antibiotics (Maheswaran et al. 1992), while that of strain EHA105 was difficult (Terakami et al. 2007). We compared 250 and 500 mg L−1 cefotaxime sodium salts for regeneration but found no significant difference between the two conditions. In the future, other antibiotics (e.g., augmentin and timentin etc.) and their concentrations should be tested for more efficient regeneration and transformation.
Another important aspect of genetic transformation is the expression level of foreign genes. Since promoters regulated gene expression both quantitatively and qualitatively, we evaluated the expression of the gusA marker gene in T. cinerariifolium for four types of promoters: the 35S promoter, the 35S promoter combined with an enhancer of El2Ω or E7Ω (Mitsuhara et al. 1996), and the bi-directional mas1'-2' promoter. The 35S promoter functions in many organs in a large number of higher plants (Benfey and Chua 1990; Terada and Shimamoto 1990; Yang and Christou 1990). However, in some species, specific promoters are required to control the high expression of the transgene. In comparisons with the 35S promoter in pBI121, the bi-directional mas1'-2' promoter was fivefold stronger, while El2Ω and E7Ω (Mitsuhara et al. 1996) were approximately fourfold stronger in terms of enhancements in GUS activity levels. Using the bi-directional mas1'-2' promoter, we succeeded in creating florists’ chrysanthemum (Chrysanthemum morifolium) with high insect and disease resistance (Shinoyama et al. 2015), and with male and female sterility (Shinoyama et al. 2020). Therefore, the bi-directional mas1'-2' promoter is expected to increase the synthesis of foreign gene products in transgenic T. cinerariifolium and florists’ chrysanthemums.
A challenge associated with genetic transformation technology is the emergence of chimaeras, which has been reported for florists’ chrysanthemums (Pavingerova et al. 1994; Benetka and Pavingerova 1995). Shinoyama et al. (2002) demonstrated that regeneration technology via calli suppressed the emergence of chimaeras in florists’ chrysanthemum. The regenerated transformed plants obtained in the present study also showed the whole-body expression of the gusA gene (see Fig. 6c), which indicates the minimized formation of chimeras.
The transformation efficiencies reported so far for T. cinerariifolium by Mao et al. (2013) and Li et al. (2022a) were 6.7% and 0.83%, respectively, when investigated using R. radiobacter. In our case, the transformation efficiency was 3.1 to 4.1% in leaf segments. Each T. cinerariifolium line exhibits a distinct genotype with large variations by its self-incompatibility (Bhat and Menary 1986; Parlevliet 1974; Teixeira da Silva 2003), which may account for the differences in transformation efficiency among these three studies. To improve transformation efficiency, the regeneration frequency from leaf segments should be increased, as well as the selection substances (e.g., paromomycin) and their concentrations should be re-examined in the future.
In Asteraceae including T. cinerariifolium, genetic transformation techniques have been established in several species. In the florists’ chrysanthemum (C. morifolium), genetically modified traits with insect and disease resistance (Takatsu et al. 1999; Toguri et al. 2003; Shinoyama et al. 2015) and unique flower colors (Noda et al. 2017) have already been produced. Genome editing technology has recently been used to create new traits in florists’ chrysanthemums (Kishi-Kaboshi et al. 2017; Shinoyama et al. 2020). Notably, Gupta et al. (2022) reported that the chrysanthemyl diphosphate synthase gene was overexpressed under the promoter CaMV35S to enhance the production of pyrethrins in Tagetes erecta, and the level of pyrethrin production in the transformants was increased to about 26-fold of that observed in non-transformed Tagetes plants. However, modifying T. cinerariifolium for enhanced pyrethrin biosynthesis is valuable because the pyrethrin production capability of this species is the highest in Asteraceae.
Pyrethrins have been reported to insecticidal effect against thrips by Yang et al. (2012). Since many cultivated crops are damaged by thrips, increased biosynthesis of pyrethrins leads to their commercial use as biopesticides for crop protection. In the present study, we have established a method for the efficient shoot regeneration and genetic transformation of T. cinerariifolium genotype #14 and confirmed the introduction and expression of the gusA gene in the T0 generation. Even though it is necessary in the future to examine seed fertility, seed setting and any other phenotypic anomaly in the T1 progeny, we believe that it will be possible to create novel T. cinerariifolium traits with increased pyrethrin production using the transformation technique established here. Also, our technique can be deployed to induce null segregants with genome editing technology to resolve the environmental safety issue.
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Aoki S, Kaneto K, Hashimoto S, Oogai H (1976) Production of pyrethrin by tissue culture. Jpn Appl Sho 51:97868 (Patent 78:24097, March, 1978)
Benetka V, Pavingerova D (1995) Phenotypic differences in transgenic plants of chrysanthemum. Plant Breed 114:169–173. https://doi.org/10.1111/j.1439-0523.1995.tb00784.x
Benfey PN, Chua NH (1990) The cauliflower mosaic virus 35S promoter: combinational regulation of transcription in plants. Sci 250:959–966. https://www.science.org/doi/10.1126/science.250.4983.959. Accessed 1 Feb 2024
Bevan MW, Flavell RB, Chilton MD (1983) A chimeric antibiotic resistance gene as a selectable marker for plant cell transformation. Nature 304:184–187. https://doi.org/10.1038/304184a0
Bhat BK, Menary RC (1986) Genotypic and phenotypic correlation in Pyrethrum ( Chrysanthemum cinerariaefolium Vis) and their implication in selection. Pyrethrum Post 16:61–65
Cashyap MM, Kueh JSH, MacKenzie IA, Pattenden G (1978) In vitro synthesis of pyrethrins from tissue cultures of Tanacetum cinerariaefolium. Phytochem 17:544–555. https://doi.org/10.1016/S0031-9422(00)89357-7
Ditta GS, Stanfield S, Corbin D, Helinski DR (1980) Broad host range DNA cloning system for Gram-negative bacteria: Construction of a gene bank of Rhizobium meliloti. Proc Natl Acad Sci USA 77:7347–7351. https://www.pnas.org/doi/abs/10.1073/pnas.77.12.7347. Accessed 1 Feb 2024
Frey L, Janick J (1991) Organogenesis in carnation. J Amer Soc Hortic Sci 116:1108–1112. https://doi.org/10.21273/JASHS.116.6.1108
Gamborg OL, Miller RA, Ojima K (1968) Nutrient requirements of suspension cultures of soybean root cells. Exp Cell Res 50:151–158. https://doi.org/10.1016/0014-4827(68)90403-5
Gupta V, Khan S, Verma RK, Shanker K, Singh SV, Rahman LU (2022) Overexpression of chrysanthemyl diphosphate synthase (CDS) gene in Tagetes erecta leads to the overproduction of pyrethrin.Transgenic Res (6):625-635. https://link.springer.com/article/10.1007/s11248-022-00323-9
Han YZ (2008) Studies on polyphenol oxidase (PPO) of compositae plant. Dissertation, Kagoshima University. http://hdl.handle.net/10232/4737. Accessed 1 Sept 2023
Hedayat M, Abdi Gh, Khosh-Khui M (2009) Regeneration via direct organogenesis from leaf and petiole segments of Pyrethrum [ Tanacetum cinerariifolium (Trevir.) Schultz-Bip.]. American-Eurasian J Agric Environ Sci 6:81–87
Hitmi A, Barthomeuf C, Sallanon H (1999) Rapid mass propagation of Chrysanthemum cinerariaefolium Vis. by callus culture and ability to synthesise pyrethrins. Plant Cell Rep 19:156–160. https://doi.org/10.1007/s002990050726
Hoekema A, Hirsch PR, Hooykaas PJJ, Schilperoort RA (1983) A binary plant vector strategy based on separation of vir- and T-region of the Agrobacterium tumefaciens Ti-plasmid. Nature 303:179–180. https://doi.org/10.1038/303179a0
Honda H, Hirai A (1990) A simple and efficient method for identification of hybrids using nonradioactive rDNA as probe. Jpn J Breed 40:339–348. https://doi.org/10.1270/jsbbs1951.40.339
Hood EE, Gelvin SB, Melchers LS, Hoekema A (1993) New Agrobacterium helper plasmids for gene transfer to plants. Transgenic Res 2:208–218. https://doi.org/10.1007/BF01977351
Jefferson RA (1987) Assay chimeric gene in plants: the GUS gene fusion system. Plant Mol Biol Rep 5:387–407. https://doi.org/10.1007/BF02667740
Kishi-Kaboshi M, Aida R, Sasaki K (2017) Generation of gene-edited Chrysanthemum morifolium using multicopy transgenes as targets and markers. Plant Cell Physiol 58:216–226. https://doi.org/10.1093/pcp/pcw222
Kosugi S, Ohashi Y, Nakajima K, Arai Y (1990) An improved assay for β-glucuronidase in transformed cells: Methanol almost completely suppresses a putative endogenous-glucuronidase activity. Plant Sci 70:133–140. https://doi.org/10.1016/0168-9452(90)90042-M
Li J, Xu Z, Zeng T, Zhou L, Li J, Hu H, Luo J, Wang C (2022a) Overexpression of TcCHS increases pyrethrin content when using a genotype-independent transformation system in Pyrethrum (Tanacetum cinerariifolium). Plants 11:1575. https://doi.org/10.3390/plants11121575
Li J, Zeng T, Xu Z, Li J, Hu H, Yu Q, Zhou L, Zheng R, Luo J, Wang C (2022b) Ribozyme-mediated CRISPR/Cas9 gene editing in pyrethrum (Tanacetum cinerariifolium) hairy roots using a RNA polymerase 2-dependent promoter. Plant Met 18:32. https://doi.org/10.1186/s13007-022-00863-5
Liu F, Wang Q, Xu P, Andreazza F, Valbon WR, Bandason E, Chen M, Yan R, Feng B, Smith LB, Scott JG, Takamatsu G, Ihara M, Matsuda K, Klimavicz J, Coats J, Oliveira EE, Du Y, Dong K (2021) A dual-target molecular mechanism of pyrethrum repellency against mosquitoes. Nat Commun 12:2553. https://doi.org/10.1038/s41467-021-22847-0
Lybrand DB, Xu H, Last RL, Pichersky E (2020) How plants synthesize pyrethrins: Safe and biodegradable insecticides. Trends Plant Sci 25:1240–1251. https://doi.org/10.1016/j.tplants.2020.06.012
Maheswaran GM, Welander M, Hutchinson JF, Graham MW, Richards D (1992) Transformation of apple rootstock M26 with Agrobacterium tumefaciens. J Plant Physiol 139:560–568. https://doi.org/10.1016/S0176-1617(11)80370-6
Manickavasagam M, Ganapathi A, Anbazhagan VR, Sudhakar B, Selvaraj N, Vasudevan A, Kasthurirengan S (2004) Agrobacterium mediated genetic transformation and development of herbicide-resistant sugarcane (Saccharum species hybrids) using axillary buds. Plant Cell Rep 23:134–143. https://link.springer.com/article/10.1007/s00299-004-0794-y. Accessed 1 Sept 2023
Mao J, Cao LY, Kong LF, Jongsma MA, Wang CY (2013) An Agrobacterium-mediated transformation system of pyrethrum (Tanacetum cinerariifolium) based on leaf explants. Sci Hortic 150:130134. https://doi.org/10.1016/j.scienta.2012.10.019
Matsuda K, Kikuta Y, Haba A, Nakayama K, Katsuda Y, Hatanaka A, Komai K (2005) Biosynthesis of pyrethrin I in seedlings of Chrysanthemum cinerariaefolium. Phytochem 66(13):1529–1535. https://doi.org/10.1016/j.phytochem.2005.05.005
Messeuger J, Arconada MC, Mele E (1993) Adventitious shoot regeneration in carnation (Dianthus caryophyllus L.). Sci Hortic 54:153–163. https://doi.org/10.1093/oxfordjournals.aob.a088097
Mitsuhara I, Ugaki M, Hirochika H, Ohshima M, Murakami T, Gotoh Y, Katayose Y, Nakamura S, Honkura R, Nishimiya S, Ueno K, Mochizuki A, Tanimoto H, Tsugawa H, Otsuki Y, Ohashi Y (1996) Efficient promoter cassettes for enhanced expression of foreign genes in dicotyledonous and monocotyledonous plants. Plant Cell Physiol 37:49–59. https://doi.org/10.1093/oxfordjournals.pcp.a028913
Murakami T, Ohashi Y (1992) Method for histochemical detection of GUS reporter gene expression in transgenic plants. Plant Cell Tech 4:281–286
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497. https://doi.org/10.1111/j.1399-3054.1962.tb08052.x
Nakamura T (1968) Studies on the Tannin of Fruits and Vegetables. Part III Polyphenolic compounds and phenol oxidative enzymes of edible burdock. Jpn Soc Food Sci Tech 15:199–206. https://doi.org/10.3136/nskkk1962.15.199
Narahashi T (1992) Nerve membrane Na+ channels as targets of insecticides. Trends Pharmacol Sci 13(6):236–241. https://doi.org/10.1016/0165-6147(92)90075-H
Noda N. Yoshioka S, Kishimoto S, Nakayama M, Douzono M, Tanaka Y, Aida R (2017) Generation of blue chrysanthemums by anthocyanin B-ring hydroxylation and glucosylation and its coloration mechanism. Sci Adv 3, e1602785. https://www.science.org/doi/10.1126/sciadv.1602785. Accessed 1 Feb 2024
Ooms G, Hooykaas PJJ, Veen RJMV, Beelen PV, Regensburg-Tuink TJG, Schilperoort RA (1982) Octopine Ti-plasmid deletion mutants of agrobacterium tumefaciens with emphasis on the right side of the T-region. Plasmid 7:15–29. https://doi.org/10.1016/0147-619X(82)90023-3
Panizzi L, Scarpati ML (1954) Constitution of cynarine, the active principle of the artichoke. Nature 174:1062–1063. https://doi.org/10.1186/1475-2891-3-5
Parlevliet JE (1974) The genetic variability of the yield components in the Kenyan pyrethrum populations. Euphytica 23:377–384. https://doi.org/10.1007/BF00035881
Pavingerova D, Dostal D, Biskova R, Benetka V (1994) Somatic embryogenesis and Agrobacterium-mediated transformation of chrysanthemum. Plant Sci 97:95–101. https://doi.org/10.1016/0168-9452(94)90111-2
Roest S, Bokelmann GS (1973) Vegetative propagation of Chrysanthemum cinerariaefolium in vitro. Sci Hortic 1:120–122. https://doi.org/10.1016/0304-4238(73)90012-5
Sasaki K, Yoshioka S, Aida R, Ohtsubo N (2021) Production of petaloid phenotype in the reproductive organs of compound flowerheads by the co-suppression of class-C genes in hexaploid Chrysanthemum morifolium. Planta 253:100. https://doi.org/10.1007/s00425-021-03605-4
Shinoyama H, Kazuma T, Komano M, Nomura Y, Tsuchiya T (2002) An efficient transformation system in chrysanthemum [Dendranthema × grandiflorum (Ramat.) Kitamura] for stable and non-chimeric expression of foreign genes. Plant Biotechnol 19:335–343. https://doi.org/10.5511/plantbiotechnology.19.335
Shinoyama H, Mitsuhara I, Mochizuki A, Kato K, Ichikawa H (2015) Transgenic chrysanthemum (Chrysanthemum morifolium Ramat) carrying both insect and disease resistance. Acta Hortic 1087:485–49. https://doi.org/10.17660/ActaHortic.2015.1087.66
Shinoyama H, Ichikawa I, Yokoi-Nishizawa A, Skaptsov M, Toki S (2020) Simultaneous TALEN-mediated knockout of chrysanthemum DMC1 genes confers male and female sterility. Sci Rep 10:16165. https://doi.org/10.1038/s41598-020-72356-1
Southern EM (1975) Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol 98:503–517. https://doi.org/10.1016/S0022-2836(75)80083-0
Takagi H, Tanaka Y, Tarumoto I, Murata N (1993) Evaluation of genetic diversity of sweet potato germplasm. I. Characterization by restriction polymorphisms analysis. Jpn J Breed 43(1):192
Takatsu Y, Nishizawa Y, Hibi T, Akutsu K (1999) Transgenic chrysanthemum (Dendranthema grandiflorum (Ramat.) Kitamura) expressing a rice chitinase gene shows enhanced resistance to gray mold (Botrytis cinerea). Sci Hort 82:113–123. https://doi.org/10.1016/S0304-4238(99)00034-5
Teixeira da Silva JA (2003) Anthemideae: advances in tissue culture, genetics and transgenic biotechnology. Afr J Biotech 2(12):547–556. https://doi.org/10.5897/AJB2003.000-1106
Terada R, Shimamoto K (1990) Expression of CaMV 35S-GUS gene in transgenic rice plants. Mol Gen Genet 220:389–392. https://doi.org/10.1007/BF00391743
Terakami S, Matsuta N, Yamamoto T, Sugaya S, Gemma H, Soejima J (2007) Agrobacterium-mediated transformation of the dwarf pomegranate (Punica granatum L.). Plant Cell Rep 26:1243–1251. https://doi.org/10.1007/s00299-007-0347-2
Toguri T, Ogawa T, Kakitani M, Tukahara M, Yoshioka M (2003) Agrobacterium-mediated transformation of chrysanthemum (Dendranthema grandiflora) plants with a disease resistance gene (pac1). Plant Biotechnol 20:121–127. https://doi.org/10.5511/plantbiotechnology.20.121
Varga F, Liber Z, Jakse J, Turudic A, Satovic Z, Radosavljevic I, Jeran N, Grdisa M (2022) Development of Microsatellite Markers for Tanacetum cinerariifolium (Trevis.) Sch. Bip., a Plant with a Large and Highly Repetitive Genome. Plants 11(13):1778. https://doi.org/10.3390/plants11131778
Wang Q, Xu P, Andreazza F, Liu Y, Nomura Y, Duran P, Jiang L, Chen M, Takamatsu G, Ihara M, Matsuda K, Isaacs R, Oliveira EE, Du Y, Dong K (2021) Identification of multiple odorant receptors essential for pyrethrum repellency in Drosophila melanogaster. PLoS Genet 17:e1009677. https://doi.org/10.1371/journal.pgen.1009677
White PR (1963) The cultivation of animal and plant cells, 2nd edn. Ronald Press Co., New York
Yamashiro T, Shiraishi A, Satake H, Nakayama K (2019) Draft genome of Tanacetum cinerariifolium, the natural source of mosquito coil. Sci Rep 9:18249. https://doi.org/10.1038/s41598-019-54815-6
Yang NS, Christou P (1990) Cell type specific expression of a CaMV 35S-GUS gene in transgenic soybean plants. Dev Genet 11:289–293. https://doi.org/10.1002/dvg.1020110407
Yang T, Stoopen G, Wiegers G, Mao J, Wang C, Dicke M, Jongsma MA (2012) Pyrethrins protect pyrethrum leaves against attack by western flower thrips, Frankliniella occidentali. J Chem Ecol 38(4):370–377. https://link.springer.com/article/10.1007/s10886-012-0097-7
Yu GR, Liu Y, Du WP, Song J, Lin M, Xu LY, Xiao FM, Liu YS (2013) Optimization of Agrobacterium tumefaciens-Mediated Immature Embryo Transformation System and Transformation of Glyphosate-Resistant Gene 2mG2-EPSPS in Maize (Zea mays L.). J Integr Agric 12:213–2142. https://doi.org/10.1016/S2095-3119(13)60567-5
Acknowledgements
We give special thanks to Drs. Shizuka Ohki (Kyoto Gakuen University), Hiroshi Kamada (University of Tsukuba), Fumio Kikuchi (Tokyo University of Agriculture), and Youichi Morinaka (Fukui Prefectural University) for their useful comments and helpful advice. Rhizobium strain EHA105 was provided by Dr. Elizabeth E. Hood. The binary vector pBIK201G was supplied by Dr. Hiroaki Ichikawa (National Institute of Agrobiological Sciences). pBE2113-GUS and pBE7133-GUS were supplied by Dr. Ichiro Mitsuhara (National Institute of Agrobiological Sciences). We also thank Ms. Ricca Murai and Chiyoko Amaya (Fukui Agricultural Experiment Station) for their technical support throughout the study. We would like to express our deep and sincere gratitude to all members at Fukui Prefectural University.
Funding
This work was supported in part by a Grant-in-Aid for Scientific Research from Fukui Prefectural University to H. S. and M. S. and the Agricultural Technology and Innovation Research Institute of Kindai University to M. H. and K. M.
Author information
Authors and Affiliations
Contributions
K.M., M.H. and H.S. designed the research, performed experiments, and analyzed data together with M.S.; H.S. and M.S. conceived the experiments. The first draft of the manuscript was written by H.S., M.S., and K.M. and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
There is no conflict of interest to declare in this study.
Financial competing interests
The authors have no financial competing interests as defined by Plant Cell, Tissue & Organ Culture, or other interests that may be perceived to influence the interpretation of the article.
Non-financial competing interests
The authors have no non-financial competing interests as defined by Plant Cell, Tissue & Organ Culture, or other interests that may be preceived to influence the interpretation of the article.
Additional information
Communicated by Goetz Hensel
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shinoyama, H., Shimizu, M., Hosokawa, M. et al. Establishment of an efficient genetic transformation system for Tanacetum cinerariifolium. Plant Cell Tiss Organ Cult 156, 97 (2024). https://doi.org/10.1007/s11240-024-02712-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11240-024-02712-w