Abstract
Invasion of alien species is one of the major environmental problems in the globalised world. Therefore, identifying pathways for the introduction and spread of alien species can help to mitigate their impact on invaded ecosystems. This study focused on the freshwater snail Gyraulus chinensis, which invades Europe from eastern Eurasia, where it is native. It was first recorded in Europe about 50 years ago and has since expanded its distribution. However, its origin and the mechanisms of its spread have not yet been studied. Therefore, we performed molecular identification and estimated its origin and dispersal mechanisms based on a literature review, molecular phylogenetics, and population genetics. We found that the haplotypes of G. chinensis in Europe are closely related to southern East Asia, namely Hong Kong, Taiwan, and the Ryukyu Islands. The results also suggest multiple introductions, mainly due to the diversity of haplotypes detected in European G. chinensis populations. Currently, the distribution and invasion of the species in the outdoor environments are concentrated in southern Europe, probably due to thermal constraints. However, the species has been recorded in many indoor habitats in the northern part of Europe, indicating a high potential for its spread in the warmer world.
Similar content being viewed by others
Introduction
Alien species cause many ecological and social problems, such as extinction of native species, degradation of agricultural land, and threats to people during outdoor activities (Pyšek et al., 2020; Cuthbert et al., 2022). In particular, freshwater environments, which are under severe threat from human activities (Vörösmarty et al., 2010), are highly affected by invasive species (Dextrase & Mandrak, 2006; Dudgeon et al., 2006; Reid et al., 2019). For example, among the 100 of the World's Worst Invasive Alien Species, about 15% are species associated with freshwater, such as Cyprinus carpio Linnaeus, 1758 and Dreissena polymorpha (Pallas, 1771) (ISSG, 2022). Considering the potential impact of alien species, it is therefore necessary to clarify the process of introduction and spread of alien species in freshwaters in terms of conservation.
Here we focused on the invasion of Gyraulus chinensis (Dunker, 1848) in Europe, which is a freshwater snail belonging to the family Planorbidae (Gastropoda: Hygrophila). This planorbid snail is native to East-Southeast Asia, although its exact range is unclear due to taxonomic inadequacies (Meier-Brook, 1983; Saito et al., 2018b, 2020). It differs from the other European representatives of the genus by having distinct black dots on a distant part of the mantle (Brown et al. 1998; Glöer, 2019; Rowson et al., 2021). The earliest documented record in Europe is from Camargue, France, in 1973 (Brown et al., 1998). Subsequently, this snail is reported to have spread over a large area in Europe, and there are also records from Africa, Australia, Middle East (Israel), and Caribbean (Martinique) as an alien species (Brown et al., 1998; Pointier, 2008; Appleton & Miranda, 2015; Mienis et al., 2015; Ponder et al., 2022). On the other hand, despite its global distribution, little is known about the nature of G. chinensis as an invasive species. At least in its native range, the snail serves as an intermediate host for parasites that are harmful to human health (Lo, 1995; Toledo et al., 1998). Although G. chinensis was a part of the community that has replaced native fauna in the Arno River in Italy (Haubrock et al., 2021), its direct impact has not been considered. Given its widespread invasion in Europe, G. chinensis may pose an existing and/or potential threat to snails of the same genus and family, which include endangered species (IUCN, 2022). In other representatives of Hygrophila, negative impacts on other native species of the same family (Lymnaeidae; Pseudosuccinea columella (Say, 1817); Vignoles et al., 2018) and the potential for introgressive hybridisation with closely related species have been reported (e.g. Ferrissia californica (Rowell, 1863); Saito et al., 2018a).
In this study, we address the clarification of the introduction and spread of G. chinensis in Europe using literature information and molecular approaches. The synthesis of literature information will trace the process of invasion in Europe and provide a better understanding of the spread process. In addition, the molecular approach will help validate the invasions of members of the genus Gyraulus, which are sometimes difficult to identify from shell morphology alone and cause taxonomic confusion (Meier-Brook, 1983; Lorencová et al., 2021), and provide insights into the origin of the introduction. By combining these approaches, this study aims to reconstruct the history of the invasion of G. chinensis. This reconstruction will not only address the further spread and potential impacts of this species, but will also expand our knowledge of the spread of alien freshwater invertebrates by comparing the history with the invasion processes of other species.
Methods
Literature survey
Some of the records of Gyraulus chinensis in Europe were documented in local malacological journals and illustrated books, which are not collected in online databases, such as Web of Science and Google Scholar. Therefore, in order to obtain as many records as possible in this study, we manually searched the scientific literature and integrated it into a list after validation (Online Resource 1). Water bodies that exist outdoors and have no artificial temperature control were treated as outdoor environments, while other water bodies, such as aquaria and heated ponds, were treated indoors. However, it cannot be completely ruled out that the sites where G. chinensis was detected were under the influence of hot water runoff, as water temperature was not documented in most surveys. In addition, some records that were not adequately documented to be outdoors were treated as possibly outdoor habitat (labelled “possibly outdoors” in Online Resource 1). The observed locations were narrowed down as much as possible using all available information. We then mapped the records using QGIS 3.16 (QGIS Development Team, 2020). In cases where it was not possible to determine the year of record, the latest possible year was used for mapping for simplicity.
Taxon sampling and DNA methods
We assembled all potential specimens of Gyraulus chinensis from our samples collected in the course of earlier investigations in outdoor freshwater environments in Albania and Italy, and indoor habitat in the Czech Republic (Fig. 1; Tables 1; Online Resource 2). To select these specimens, we followed morphological characters of G. chinensis reported in the literature (Brown et al. 1998; Glöer, 2019; Rowson et al., 2021). Some of the Italian specimens were collected in the 1990s and early 2000s. In addition, the Albanian populations were the first records. For comparison, we also collected and analysed G. chinensis specimens from native Asian regions, including Hong Kong, near the type locality of G. chinensis, and the other six Gyraulus species (Tables 1; Online Resource 2). Some of these samples were used in two previous studies, and fixation and extraction were performed using the methods described in the respective study (Saito et al., 2018a, b; Lorencová et al., 2021). Other newly collected specimens were fixed in 96% ethanol or dried at ambient temperature and humidity. Total DNA was extracted using the E.Z.N.A. Mollusc DNA Kit (Omega BioTek, GA, USA) according to the manufacturer's instructions.
We then amplified two mitochondrial regions, cytochrome b (CytB) and cytochrome c oxidase subunit I (CO1), and two nuclear genes, internal transcribed spacers 1 (ITS1) and 2 (ITS2) of the rRNA gene cassette. The primers and protocols for amplification were the same as in Lorencová et al. (2021). Each amplified product was purified with ExoSAP (Affymetrix, CA, USA) and sequenced at the OMICS Core facility, BIOCEV (Czech Republic) using the BigDye® Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, CA, USA) or SEQme s.r.o. (Czech Republic) using the Gerbera Sequencing Kit v3.1 (SEQme, Czech Republic). After removing the primer sequences, all sequences were deposited in GenBank (Accession Nos OQ121085–OQ121110, OQ127675–OQ127702, OQ133368–OQ133396, and OQ133368–OQ133396).
Molecular phylogenetics and population genetics
Phylogenetic analyses for molecular identification of the introduced Gyraulus chinensis, which taxonomy is insufficient, were performed using two mitochondrial (CytB and CO1) and two nuclear regions (ITS1 and ITS2). Sequences of six Gyraulus species from GenBank, validated in Lorencová et al. (2021), were used for comparison. The resulting CO1 + CytB dataset included 39 individuals with 78 sequences (25 of which were from GenBank), 33 for ITS1 (10 from GenBank) and 39 for ITS2 (10 from GenBank) (Table 1). Phylogenetic analyses had to be conducted separately for ITS1 and ITS2 regions because we were not able to amplify both markers for some specimens. In addition, with reference to Lorencová et al. (2021), Planorbis planorbis (Linnaeus, 1758) was used as an obvious outgroup. Each dataset was aligned using MUSCLE (Edgar, 2004) implemented in MEGA 7 (Kumar et al., 2016) with default settings. To reduce uncertainty for phylogenetic analyses, gaps in the alignments of CO1, ITS1 and ITS2 were eliminated using trimAl 1.2 (Capella-Gutiérrez et al., 2009). Finally, identical haplotypes were stacked with FaBox 1.61 (Villesen, 2007).
Maximum likelihood analyses (ML) were performed using W- IQ-TREE (Nguyen et al., 2015; Trifinopoulos et al., 2016) with the ‘-spp’ option (same set of branch lengths in all partitions, allowing each partition to evolve at its own rate). Evolutionary models and partitions for ML analyses were selected using PartitionFinder 2 (Lanfear et al., 2016) based on AICc (Table 2). The ML tree topologies were evaluated using ultrafast bootstrapping (UFB; Hoang et al., 2018) with 10,000 replicates, Shimodaira-Hasegawa approximate likelihood ratio test (SH-aLRT; Kishino & Hasegawa, 1989; Shimodaira & Hasegawa, 1999) with 10,000 replicates, and a Bayes-like transformation of aLRT (aBayes; Anisimova et al., 2011). Bayesian inference analyses (BI) were performed with MrBayes 3.2.7a (Ronquist & Huelsenbeck, 2003) using four simultaneous chains and sampling trees every 1000 generations. The ‘usebeagle’ option was turned off to avoid unexpected errors. Four chains were run for 1 million generations in all datasets. After using Tracer v. 1.7 (Rambaut et al., 2018) to confirm that the effective sample size (ESS) of all parameters in each dataset was greater than 200, we discarded the first 101 trees as burn-in after checking for convergence. The remaining trees were then used to estimate each tree with maximum clade reliability and to evaluate each tree topology with Bayesian posterior probability (BPP).
To reveal the origin(s) and invasion history of G. chinensis in Europe, we explored the genetic structure of the G. chinensis populations using the CO1 region, which has the largest number of relevant sequences in GenBank. However, the taxonomy of the genus Gyraulus is not fully clarified, and it is difficult to distinguish G. chinensis a priori based on the GenBank registry name. Therefore, after obtaining all Gyraulus sequences from GenBank, we performed preliminary phylogenetic analyses and selected sequences that fell into the monophyletic clade that was sister to G. parvus (Say, 1817). This clade also contained sequences previously considered to represent G. chinensis and also those from near to the type locality. Consequently, 164 haplotypes from 284 sequences, including sequences from Hong Kong, close to the type locality of G. chinensis, were used for the subsequent analysis (Online Resource 2). In addition, two sequences of G. parvus were selected as outgroups. Phylogenetic analysis was then performed using this CO1 dataset. After alignment, the sequences were stacked by haplotype, and then the ML and BI analyses were performed in the same flow as the previous analyses (the evolutionary models in these analyses are listed in Table 2). However, the number of generations in the BI analysis was 3 million and the burn-in was the first 301 trees. Finally, we presented a haplotype network using the monophyly in the CO1 phylogeny for G. chinensis and included all haplotypes detected in Europe. The haplotype network was constructed with the TCS algorithm (Clement et al., 2000) using PopART (Leigh & Bryant, 2015).
Results
Distribution of Gyraulus chinensis
Based on the collected 59 literature records and our survey, Gyraulus chinensis has been reported outdoors from Albania, Austria, France, Germany, Greece, Italy, Montenegro, the Netherlands, Portugal and Spain, and indoors from Austria, the Czech Republic, Ireland, Germany, the Netherlands, Norway, Sweden and the United Kingdom (Fig. 1; Online Resource 1). In Belgium, the occurrence was highly speculated (Van Goethem, 1992; Sablon & Vercauteren, 2011), but no evidence was found in the literature during our study. The south of Wonseradeel in the Netherlands was the northern limit of occurrence of G. chinensis in the outdoor environments.
After the first outdoor record from France (1973), the species was also reported outdoors from Italy, Portugal and Spain by the 1980s. Findings in other European regions are more recent. In south-eastern Europe, the species was first reported outdoors from Greece in 2003, from Montenegro in 2013 and from Albania in 2018. In Central Europe, the oldest outdoor records are from the Netherlands (1976). In Austria and Germany, possible occurrences in the outdoor environments were reported already in 1993 and 1998, but reliable outdoor records come from 2001 and 2009. The oldest records from indoor habitats are from Ireland and the United Kingdom (1974), and from Austria and the Netherlands as early as the 1990s. In other countries, it has been recorded indoors since the 2000s. In France, Spain, Italy, and south-eastern Europe, there are no documented records in an explicitly indoor habitat.
Molecular identification
Our three (CO1 + CytB, ITS1 and ITS2) phylogenies showed that most Gyraulus species formed monophylies that were well supported (> 90 and > 0.95) by at least two evaluation values (Fig. 2; Online Resource 3). Gyraulus chinensis in Europe has nine haplotypes in CO1 + CytB, eleven haplotypes in ITS1 and five haplotypes in ITS2, although some European samples were not sequenced. Samples from Albania, Italy, and Japan shared one haplotype in ITS2. In three phylogenies, the European G. chinensis haplotypes were included in the well-supported (CO1 + CytB: UFB = 99/ SH -aLRT = 95/aBayes = 1.00/BPP = 1.00; ITS1: 100/100/1.00/1.00; ITS2: 91/96/1.00/1.00) monophyly of G. chinensis with samples from Japan, Taiwan, and Hong Kong (Fig. 2; Online Resource 3). This is an important outcome as Hong Kong sequences represent data from close to the type locality.
Maximum likelihood phylogenies of the genus Gyraulus inferred from nuclear internal transcribed spacer 2 (ITS2), and mitochondrial cytochrome b (Cytb) and cytochrome c oxidase subunit 1 (CO1) DNAs. The numbers at each node indicate support values by four different methods. The labels on each haplotype represent the IDs of the included samples (Table 1). The bars next to the labels indicate the species based on a previous molecular phylogeny (Lorencová et al. 2021). The colours of each ID represent the countries or regions collected
Population genetic structure
Our CO1 phylogeny revealed the phylogenetic relationship of Gyraulus chinensis and closely related lineages (Fig. 3). Many basal nodes were not supported, while most terminal nodes were well supported and often consisted of geographically related haplotypes. All European G. chinensis were included in one relatively well-supported monophyly (91/97/1.00/-), and in addition, the remaining haplotypes from Europe, with the exception of one haplotype from Italy, were included in another well-supported monophyly (100/100/1.00/-).
Maximum likelihood phylogeny of Gyraulus chinensis and closely related lineages inferred from mitochondrial cytochrome c oxidase subunit 1 (CO1) DNA. The numbers at each node indicate the support values by four different methods (they are not shown at the terminal nodes due to visibility). The label on each operational unit represents the haplotype numbers (Online Resource 2), and for European samples the country where the haplotype was collected. The colours of each label represent the countries or geographical areas, which are shown on the map at the top left. The geographical areas have been defined with reference to biogeographical boundaries for convenience. This map was created from the Global Self-consistent, Hierarchical, High-resolution Geography Database, version 2.3.5 (Wessel & Smith 1996, 2016), using QGIS version 3.16 (QGIS Development Team, 2020). The sites were identified based on the publications and GenBank registry information of each sequence (Online Resource 2)
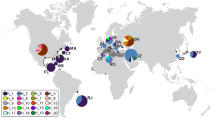
The haplotype network, illustrated by the monophyly with all nine European haplotypes, showed the geographical relationship between the individual haplotypes (Fig. 4; Table 2). Haplotype No. 1 included eight sequences from Albania and one sequence from the Ryukyu Islands, Japan, and Taiwan. All other European sequences were included in eight haplotypes that did not include sequences from outside Europe. Each haplotype, with the exception of haplotype No. 7 from Italy, had the most closely related haplotypes containing sequence(s) from the southern part of East Asia, namely Taiwan, Hong Kong, and the Ryukyu Islands of Japan. In addition, haplotype No. 7 consisted of two Italian sequences from Arenzano and Venaria, and the most closely related haplotype was from Kyushu Island in Japan.
The TCS network of mitochondrial CO1 haplotypes for the monophyly from the phylogeny in Fig. 3. The numbers with circles represent the nominal haplotype numbers (Online Resource 2), and each circle size represents the haplotype frequency. The bar on the branch indicates a mutation. Each colour represents countries or geographical areas shown on the map in the upper right part. This map was created from the Global Self-consistent, Hierarchical, High-resolution Geography Database, version 2.3.5 (Wessel & Smith, 1996, 2016), using QGIS version 3.16 (QGIS Development Team, 2020)
Discussion
Origins of Gyraulus chinensis invasion
This is the first study to demonstrate the introduction of Gyraulus chinensis into Europe based on extensive molecular phylogenies, as previously this was only supported based on morphological identification (Figs. 2; Online Resource 3). Although there have been some previous sequences from Montenegro (von Oheimb et al., 2013; Saito et al., 2022) and British aquaria (Rowson et al. 2021) assumed to represent G. chinensis, they were identified as Gyraulus sp. Therefore, our study provides the first indisputable evidence for G. chinensis occurrence in Europe based on sequences from several European countries. Moreover, our CO1 phylogeny and haplotype network revealed a strong genetic relationship between G. chinensis from Europe and from Hong Kong (New Territories), close to the type locality of G. chinensis (Hong Kong Island; Dunker, 1848; Figs. 3, 4). Therefore, despite the confusion in the taxonomy of G. chinensis, the European G. chinensis collected in this study should strictly be G. chinensis. However, because our sampling may not have captured the entire European diversity of G. chinensis populations, it cannot be ruled out that there are more lineages of G. chinensis sensu lato introduced into Europe than those already detected. Therefore, further sampling and taxonomic studies of G. chinensis and related species are needed to clarify this issue.
Based on our molecular results, it is difficult to fully determine the origin of introduced G. chinensis in Europe, because the GenBank sequences of G. chinensis and related lineages, although relatively extensive, could not cover all native distributions, and CO1 sequences can be affected by introgressive hybridisation and ancestral polymorphisms (Ballard, 2000; Ballard & Whitlock, 2004). Nevertheless, the European populations may have originated in the south-eastern part of the Eurasian continent, particularly on Kyushu Island, the Ryukyu Islands, Taiwan, and Hong Kong. At a minimum, we can conclude that European haplotypes have little relationship with haplotypes from South Asia, Central Asia, Northeast Asia and North Asia.
The haplotypes of G. chinensis in Europe were diverse, with haplotype No. 7 from Italy being genetically quite distant. Moreover, the localities of the first outdoor records are geographically distant and dated over a long period of time (Fig. 1; Online Resource 1). These findings suggest the possibility that introduction occurred from more than one source or, less likely, from a source that exhibited high haplotype diversity. Most records of G. chinensis in the outdoor environments are from southern Europe, and date from the early stages of the invasion, while records from the north are relatively recent and rare, as suggested by our field and literature surveys (Fig. 1; Online Resource 1). This suggests that temperature differences between the native range and introduced areas were important in maintaining populations during the early stages of the invasion. In general, temperature can be a major factor determining the success or failure of an alien species introduction and establishment (Hayes & Barry, 2008; Petitpierre et al., 2012; Guisan et al., 2014), and it has also been suggested that niche conservatism for temperature may have influenced the formation of the distribution of Hygrophila snails (Saito et al., 2021).
Mechanisms of introduction
Although the specific mechanisms of introduction remain unclear, the most likely hypothesis is invasion via the aquarium trade. Invasion of freshwater organisms via the aquarium trade is often considered an influential mechanism (e.g. Papavlasopoulou et al., 2014; Nunes et al., 2015; Ng et al., 2016; Vinarski, 2017). In particular, small invertebrates, such as Hygrophila snails, are often imported and introduced as unintended hitchhikers of target species in the trade (Duggan, 2010; Patoka et al., 2016, 2017; Duggan & Pullan, 2017). In our case, Gyraulus chinensis is commonly found in indoor aquaria in botanical gardens (Fig. 1; Online Resource 1), and the Czech haplotypes from indoor aquaria are genetically close to the Italian and Albanian haplotypes in the outdoor environments. In addition, CO1 sequences from UK aquaria (but not published) have been reported to be identical to the sequences from Taiwan (Rowson et al., 2021). These records suggest that aquaria are important sources of G. chinensis invasion in Europe. This hypothesis is also supported by the presumed origin(s) of European G. chinensis on the south-eastern Eurasian continent, as European countries mainly import freshwater organisms from the tropics, such as Singapore and Hong Kong (Madsen & Frandsen, 1989; Brunel, 2009). In addition, the genetic diversity of G. chinensis in Europe could be explained by the fact that the introduced populations in indoor aquaria across Europe might represent a source of potentially high genetic diversity. On the other hand, there is no evidence to rule out other mechanisms of introduction and spread. In particular, due to the wide and dense distribution of G. chinensis, it is difficult to explain all dispersal in Europe by the aquarium trade alone. Other mechanisms such as release of contaminants by fisheries, transport by humans, external and internal transport by birds, and natural dispersal via water currents should be examined in further studies. Based on the literature records (Fig. 1; Online Resource 1), natural dispersal in particular could be the major contributor to dispersal, as many records in Italy and Spain have been documented within a short period of time.
Distribution in Europe
Currently, the records of introduced Gyraulus chinensis populations are concentrated in southern Europe (Fig. 1). There are also records in the northern regions, i.e. the Netherlands, Austria, and Germany, but findings in the outdoor environments seem to be much rarer than in southern Europe (Neckheim, 2017). Moreover, most of these northern records are from urban areas and even in the outdoor environments, it cannot be excluded that they inhabit water bodies affected by hot water discharge. Indeed, in the Lower Rhine, G. chinensis only occurs in water bodies with hot water discharge (van der Velde et al., 2002). As mentioned above, this limitation of invasion might be caused by climatic differences between the native range and Europe, except for the southern parts. However, climate change and adaptation of alien species may allow them to spread to new environments (Sax et al., 2007; Walther et al., 2009; Chapman et al., 2017; Hulme, 2017). Furthermore, as G. chinensis is observed in many indoor aquaria, there is a large invasion debt (i.e. the time lag of introduction; Rouget et al., 2016) in many areas of Europe. Therefore, its spread may continue in the future under the climate warming scenario. On the other hand, the consequences of G. chinensis invasion have hardly been assessed and documented so far. The impacts of non-native species vary depending on the regions and ecosystems (Uchida et al., 2016) and may even change due to their adaptive evolution (Prentis et al., 2008; Whitney & Gabler, 2008). Therefore, it will be necessary to further assess and carefully monitor the impacts.
Data availability
All data generated during this study are included in this published article, its supplementary information files, and deposited in GenBank [OQ121085–OQ121110, OQ127675–OQ127702, OQ133368–OQ133396, and OQ158876–OQ158898].
References
Anisimova, M., M. Gil, J.-F. Dufayard, C. Dessimoz & O. Gascuel, 2011. Survey of branch support methods demonstrates accuracy, power, and robustness of fast likelihood-based approximation schemes. Systematic Biology 60: 685–699.
Appleton, C. C. & N. A. F. Miranda, 2015. Two Asian freshwater snails newly introduced into South Africa and an analysis of alien species reported to date. African Invertebrates 56: 1–17.
Ballard, J. W. O., 2000. When one is not enough: Introgression of mitochondrial DNA in Drosophila. Molecular Biology and Evolution 17: 1126–1130.
Ballard, J. W. O. & M. C. Whitlock, 2004. The incomplete natural history of mitochondria. Molecular Ecology 13: 729–744.
Brown, D. S., M. A. A. Gracio & C. Meier-Brook, 1998. The Asian freshwater snail Gyraulus chinensis (Dunker, 1848) (Planorbidae) in West Africa and Europe. Journal of African Zoology 112: 203–213.
Brunel, S., 2009. Pathway analysis: aquatic plants imported in 10 EPPO countries. EPPO Bulletin 39: 201–213.
Capella-Gutiérrez, S., J. M. Silla-Martinez & T. Gabaldon, 2009. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25: 1972–1973.
Chapman, D. S., R. Scalone, E. Štefanić & J. M. Bullock, 2017. Mechanistic species distribution modeling reveals a niche shift during invasion. Ecology 98: 1671–1680.
Clement, M., D. Posada & K. A. Crandall, 2000. TCS: a computer program to estimate gene genealogies. Molecular Ecology 9: 1657–1659.
Cuthbert, R. N., C. Diagne, P. J. Haubrock, A. J. Turbelin & F. Courchamp, 2022. Are the “100 of the world’s worst” invasive species also the costliest? Biological Invasions 24: 1895–1904.
Dextrase, A. J. & N. E. Mandrak, 2006. Impacts of alien invasive species on freshwater fauna at risk in Canada. Biological Invasions 8: 13–24.
Duggan, I. C., 2010. The freshwater aquarium trade as a vector for incidental invertebrate fauna. Biological Invasions 12: 3757–3770.
Dunker, W., 1848. Diagnoses specierum novarum generis Planorbis collectionis Cumingianae. Proceedings of the Zoological Society of London 16: 40–43.
Dudgeon D., A. H. Arthington, M. O. Gessner, Z.-I. Kawabata, D. J. Knowler, C. Lévêque, R. J. Naiman, A.-H. Prieur-Richard, D. Soto, M. L. J. Stiassny & C. A. Sullivan, 2006. Freshwater biodiversity: importance threats status and conservation challenges. Biological Reviews 81: 163–182.
Duggan, I. C. & S. G. Pullan, 2017. Do freshwater aquaculture facilities provide an invasion risk for zooplankton hitchhikers? Biological Invasions 19: 307–314.
Edgar, R. C., 2004. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Research 32: 1792–1797.
Glöer, P., 2019. The freshwater gastropods of the West-Palaearctis, Volume 1. Biodiversity Research Lab, Hetlingen, Germany. pp. 1–399.
Guisan, A., B. Petitpierre, O. Broennimann, C. Daehler & C. Kueffer, 2014. Unifying niche shift studies: insights from biological invasions. Trends in Ecology & Evolution 29: 260–269.
Haubrock, P. J., F. Pilotto, G. Innocenti, S. Cianfanelli & P. Haase, 2021. Two centuries for an almost complete community turnover from native to non-native species in a riverine ecosystem. Global Change Biology 27: 606–623.
Hayes, K. R. & S. C. Barry, 2008. Are there any consistent predictors of invasion success? Biological Invasions 10: 483–506.
Hoang, D. T., O. Chernomor, A. von Haeseler, B. Q. Minh & L. S. Vinh, 2018. UFBoot2: Improving the ultrafast bootstrap approximation. Molecular Biology and Evolution 35: 518–522.
Hulme, P. E., 2017. Climate change and biological invasions: evidence, expectations, and response options: alien species and climate change in Great Britain. Biological Reviews 92: 1297–1313.
IUCN., 2022. The IUCN Red List of threatened species. Version 2022-1. Retrieved from https://www.iucnredlist.org (12 August 2022).
ISSG., 2022. Global Invasive Species Database. Retrieved from http://www.iucngisd.org/gisd/100_worst.php (10 August 2022).
Kishino, H. & M. Hasegawa, 1989. Evaluation of the maximum likelihood estimate of the evolutionary tree topologies from DNA sequence data, and the branching order in hominoidea. Journal of Molecular Evolution 29: 170–179.
Kumar, S., G. Stecher & K. Tamura, 2016. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecular Biology and Evolution 33: 1870–1874.
Lanfear, R., P. B. Frandsen, A. M. Wright, T. Senfeld & B. Calcott, 2016. PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Molecular Biology and Evolution 34: 772–773.
Leigh, J. W. & D. Bryant, 2015. POPART: full-feature software for haplotype network construction. Methods in Ecology and Evolution 6: 1110–1116.
Lo, C. T., 1995. Echinostoma macrorchis: life history, population dynamics of intramolluscan stages, and the first and second intermediate hosts. The Journal of Parasitology 81: 569–576.
Lorencová, E., L. Beran, M. Nováková, V. Horsáková, B. Rowson, J. Č Hlaváč, J. C. Nekola & M. Horsák, 2021. Invasion at the population level: a story of the freshwater snails Gyraulus parvus and G. laevis. Hydrobiologia 848: 4661–4671.
Madsen, H. & F. Frandsen, 1989. The spread of freshwater snails including those of medical and veterinary importance. Acta Tropica 46: 139–146.
Meier-Brook, C., 1983. Taxonomic studies on Gyraulus (Gastropoda: Planorbidae). Malacologia 24: 1–113.
Mienis, H. K., S. Vaisman & O. Rittner, 2015. Finds of Gyraulus chinensis (Dunker, 1848) (Mollusca, Gastropoda, Planorbidae) in Israel. Triton 31: 9–11.
Neckheim, C. M., 2017. De veranderde Noord-Hollandse slakkenfauna. Tussen Duin & Dijk 16: 16–19.
Ng, T. H., S. K. Tan, W. H. Wong, R. Meier, S.-Y. Chan, H. H. Tan & D. C. J. Yeo, 2016. Molluscs for sale: assessment of freshwater gastropods and bivalves in the ornamental pet trade. PLOS ONE 11: e0161130.
Nguyen, L.-T., H. A. Schmidt, A. von Haeseler & B. Q. Minh, 2015. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Molecular Biology and Evolution 32: 268–274.
Nunes, A., E. Tricarico, V. Panov, A. Cardoso & S. Katsanevakis, 2015. Pathways and gateways of freshwater invasions in Europe. Aquatic Invasions 10: 359–370.
von Oheimb, P. V., C. Albrecht, F. Riedel, U. Bössneck, H. Zhang & T. Wilke, 2013. Testing the role of the Himalaya Mountains as a dispersal barrier in freshwater gastropods (Gyraulus spp.). Biological Journal of the Linnean Society 109: 526–534.
Papavlasopoulou, I., C. Perdikaris, L. Vardakas & I. Paschos, 2014. Enemy at the gates: introduction potential of non-indigenous freshwater crayfish in Greece via the aquarium trade. Open Life Sciences 9: 11–18.
Patoka, J., M. Bláha, M. Devetter, K. Rylková, Z. Čadková & L. Kalous, 2016. Aquarium hitchhikers: attached commensals imported with freshwater shrimps via the pet trade. Biological Invasions 18: 457–461.
Patoka, J., O. Kopecký, V. Vrabec & L. Kalous, 2017. Aquarium molluscs as a case study in risk assessment of incidental freshwater fauna. Biological Invasions 19: 2039–2046.
Petitpierre, B., C. Kueffer, O. Broennimann, C. Randin, C. Daehler & A. Guisan, 2012. Climatic niche shifts are rare among terrestrial plant invaders. Science 335: 1344–1348.
Pointier, J.-P., 2008. Guide to the freshwater molluscs of the Lesser Antilles, ConchBooks, Hackenheim, Germany:, 128.
Ponder, W. F., A. Hallan, M. Shea, S. A. Clark, K. Richards, M. W. Klunzinger, & V. Kessner, 2022. Australian freshwater molluscs. Revision 1A. Retrieved from https://keys.lucidcentral.org/keys/v3/freshwater_molluscs/ (12 August 2022).
Prentis, P. J., J. R. U. Wilson, E. E. Dormontt, D. M. Richardson & A. J. Lowe, 2008. Adaptive evolution in invasive species. Trends in Plant Science 13: 288–294.
Pyšek, P., P. E. Hulme, D. Simberloff, S. Bacher, T. M. Blackburn, J. T. Carlton, W. Dawson, F. Essl, L. C. Foxcroft, P. Genovesi, J. M. Jeschke, I. Kühn, A. M. Liebhold, N. E. Mandrak, L. A. Meyerson, A. Pauchard, J. Pergl, H. E. Roy, H. Seebens, M. Kleunen, M. Vilà, M. J. Wingfield & D. M. Richardson, 2020. Scientists’ warning on invasive alien species. Biological Reviews 95: 1511–1534.
QGIS Development Team., (2020). QGIS geographic information system Version 3.16. Open source geospatial foundation project. Retrieved from http://qgis.osgeo.org (12 Dec 2020).
Rambaut, A., A. J. Drummond, D. Xie, G. Baele & M. A. Suchard, 2018. Posterior summarization in Bayesian phylogenetics using tracer 1.7. Systematic Biology 67: 901–904.
Reid, A. J., A. K. Carlson, I. F. Creed, E. J. Eliason, P. A. Gell, P. T. J. Johnson, K. A. Kidd, T. J. MacCormack, J. D. Olden, S. J. Ormerod, J. P. Smol, W. W. Taylor, K. Tockner, J. C. Vermaire, D. Dudgeon & S. J. Cooke, 2019. Emerging threats and persistent conservation challenges for freshwater biodiversity. Biological Reviews 94: 849–873.
Ronquist, F. & J. P. Huelsenbeck, 2003. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19: 1572–1574.
Rouget, M., M. P. Robertson, J. R. U. Wilson, C. Hui, F. Essl, J. L. Renteria & D. M. Richardson, 2016. Invasion debt: quantifying future biological invasions. Diversity and Distributions 22: 445–456.
Rowson, B., H. A. Powell, M. J. Willing, M. Dobson, & H. Shaw, 2021. Freshwater snails of Britain and Ireland. Field Studies Council, Shropshire, UK. pp. 1–191.
Sablon, R. & T. Vercauteren, 2011. Exotische soorten weekdieren in (Antwerpse) rivieren en stilstaande waters. Evolutie van de voorbije 20 jaar. Antenne 5: 9–17.
Saito, T., V. T. Do, L. Prozorova, T. Hirano, H. Fukuda & S. Chiba, 2018a. Endangered freshwater limpets in Japan are actually alien invasive species. Conservation Genetics 19: 947–958.
Saito, T., T. Hirano, L. Prozorova, V. Tu Do, A. Sulikowska-Drozd, T. Sitnikova, P. Surenkhorloo, D. Yamazaki, Y. Morii, Y. Kameda, H. Fukuda & S. Chiba, 2018b. Phylogeography of freshwater planorbid snails reveals diversification patterns in Eurasian continental islands. BMC Evolutionary Biology 18: 164.
Saito, T., S. Chiba & H. Fukuda, 2020. Type materials of the species of the Planorbidae (Mollusca, Gastropoda, Hygrophila) described by Shuichi Mori. Molluscan Research 40: 169–182.
Saito, T., T. Hirano, B. Ye, L. Prozorova, M. S. Shovon, T. V. Do, K. Kimura, P. Surenkhorloo, Y. Kameda, Y. Morii, H. Fukuda & S. Chiba, 2021. A comprehensive phylogeography of the widespread pond snail genus Radix revealed restricted colonization due to niche conservatism. Ecology and Evolution 11: 18446–18459.
Saito, T., T. Sasaki, Y. Tsunamoto, S. Uchida, K. Satake, Y. Suyama, & S. Chiba, 2022. Even short distance dispersal over a barrier can affect genetic differentiation in Gyraulus, an island freshwater snail. Freshwater Biology 67: 1971–1983.
Sax, D., J. Stachowicz, J. Brown, J. Bruno, M. Dawson, S. Gaines, R. Grosberg, A. Hastings, R. Holt & M. Mayfield, 2007. Ecological and evolutionary insights from species invasions. Trends in Ecology & Evolution 22: 465–471.
Shimodaira, H. & M. Hasegawa, 1999. Multiple comparisons of log-likelihoods with applications to phylogenetic inference. Molecular Biology and Evolution 16: 1114–1116.
Toledo, R., C. Muñoz-Antolí, M. Pérez & J. G. Esteban, 1998. Larval trematode infections in freshwater gastropods from the Albufera Natural Park in Spain. Journal of Helminthology 72: 79–82.
Trifinopoulos, J., L.-T. Nguyen, A. von Haeseler & B. Q. Minh, 2016. W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Research 44: W232–W235.
Uchida, S., H. Mori, T. Kojima, K. Hayama, Y. Sakairi & S. Chiba, 2016. Effects of an invasive ant on land snails in the Ogasawara Islands: Impact of an invasive ant on land snails. Conservation Biology 30: 1330–1337.
Van Goethem, J. L., 1992. A current state of knowledge on the taxonomy and distribution of non-marine molluscs in Belgium. Faunal inventories of sites for cartography and nature conservation - Proceedings of the 8th International Colloqium of the European Invertebrate Survey: 189–194.
van der Velde, G., I. Nagelkerken, S. Rajagopal, & A. B. de Vaate, 2002. Invasions by alien species in inland freshwater bodies in Western Europe: The Rhine Delta. In Leppäkoski, E., S. Gollasch, & S. Olenin (eds), Invasive aquatic species of Europe. Distribution, impacts and management. Dordrecht, the Netherlands. pp. 360–372.
Vignoles, P., G. Dreyfuss & D. Rondelaud, 2018. Consequences of invasion by Pseudosuccinea columella on the dynamics of native lymnaeids living on the acid soils of central France. Molluscan Research 38: 287–295.
Villesen, P., 2007. FaBox: an online toolbox for fasta sequences. Molecular Ecology Notes 7: 965–968.
Vinarski, M. V., 2017. The history of an invasion: phases of the explosive spread of the physid snail Physella acuta through Europe, Transcaucasia and Central Asia. Biological Invasions 19: 1299–1314.
Vörösmarty, C. J., P. B. McIntyre, M. O. Gessner, D. Dudgeon, A. Prusevich, P. Green, S. Glidden, S. E. Bunn, C. A. Sullivan, C. R. Liermann & P. M. Davies, 2010. Global threats to human water security and river biodiversity. Nature 467: 555–561.
Walther, G.-R., A. Roques, P. E. Hulme, M. T. Sykes, P. Pyšek, I. Kühn, M. Zobel, S. Bacher, Z. Botta-Dukát & H. Bugmann, 2009. Alien species in a warmer world: risks and opportunities. Trends in Ecology & Evolution 24: 686–693.
Wessel, P. & W. H. F. Smith, 1996. A global, self-consistent, hierarchical, high-resolution shoreline database. Journal of Geophysical Research 101: 8741–8743.
Wessel, P., & W. H. F. Smith, 2016. GSHH: A global, self-consistent, hierarchical, high-resolution shoreline database, Version 2.3.5. Retrieved from http://www.soest.hawaii.edu/pwessel/gshhg/ (4 Dec 2020).
Whitney, K. D. & C. A. Gabler, 2008. Rapid evolution in introduced species, ‘invasive traits’ and recipient communities: challenges for predicting invasive potential: Evolution and invasion predictions. Diversity and Distributions 14: 569–580.
Acknowledgements
We are grateful to Marco Bodon (Natural History Museum University of Florence, Italy) for providing material from Italy.
Funding
Open access publishing supported by the National Technical Library in Prague. The study was financially supported by the Czech Science Foundation (P504/20-18827S).
Author information
Authors and Affiliations
Contributions
MH and EŠ conceived the idea. EL, LB, BY, MN, and VH generated the data. TS analysed the data and led writing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
All research pertaining to this article did not require any research permit(s).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Guest editors: Manuel P. M. Lopes-Lima, Lyubov E. Burlakova, Ting Hui Ng, Alexandra Zieritz & Ronaldo G. Sousa / Biology and Impact of Invasive Freshwater Molluscs
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Saito, T., Šlachtová, E., Nováková, M. et al. Invasion history of Gyraulus chinensis (Gastropoda: Planorbidae) in Europe: a molecular and literature-based approach. Hydrobiologia (2023). https://doi.org/10.1007/s10750-023-05157-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10750-023-05157-y