Abstract
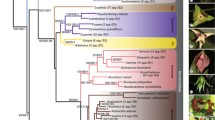
The Andes represent a hotspot of diversity within Cactaceae. Owing to its small globular growth and its diverse flower colours the Andean genus Aylostera Speg. has drawn the attention of many cactus enthusiasts. Based on molecular data, the genus was recently split from Rebutia s.l. In contrast to the large number of taxa described within Aylostera, a comprehensive phylogenetic reconstruction is still warranted. The aim of our study was to reconstruct a phylogeny of Aylostera and to review its taxonomy based on morphological characters. We sequenced the chloroplast DNA markers atpB-rbcL and trnS-trnG intergenic spacers and scored amplified fragment lengths polymorphisms (AFLPs). We reconstructed phylogenetic trees and analysed results of AFLP neighbour-nets, Bayesian clustering and Principal Coordinate Analysis based on this genetic data. Our data demonstrated that Aylostera is monophyletic and is best recognized as a separate genus because it is only distantly related to the clade containing the type species of Rebutia. Within Aylostera we detected three clades—the Digitorebutia clade, the Aylostera sensu stricto clade and the A. einsteinii clade—which are characterized by unique combinations of morphological character states. The species composition of these clades was largely congruent between the molecular markers but AFLP data indicated some admixture. Based on genetic and morphological characters, we propose a new classification of Aylostera and present a key to species recognizing nine species.







Similar content being viewed by others
References
Akaike H (1974) A new look at the statistical model identification. IEEE Trans Autom Control 19:716–723
Anderson EF (2001) The cactus family. Timber Press, Portland
Anderson EF (2005) Das große Kakteenlexikon. Eugen Ulmer, Stuttgart
Arakaki M, Christin P-A, Nyffeler R, Lendel A, Eggli U, Ogburn RM, Spriggs E, Moore MJ, Edwards EJ (2011) Contemporaneous and recent radiations of the world’s major succulent plant lineages. Proc Natl Acad Sci USA 108:8379–8384. doi:10.1073/pnas.1100628108
Backeberg C (1958–1962) Die Cactaceae. Gustav Fischer Verlag, Jena
Backeberg C (1966) Das Kakteenlexikon. Gustav Fischer Verlag, Jena
Bárcenas RT, Yesson C, Hawkins JA (2011) Molecular systematics of the Cactaceae. Cladistics 27:1–20. doi:10.1111/j.1096-0031.2011.00350.x
Barthlott W, Burstedde K, Geffert JL, Ibisch PL, Korotkova N, Miebach A, Rafiqpoo MD, Stein A, Mutke J (2015) Biogeography and biodiversity of cacti. Biogeographie und Biodiversität der Kakteen. Schumannia 7:208
Bruneau A, Starr JR, Joly S (2007) Phylogenetic relationships in the genus Rosa: new evidence from chloroplast DNA sequences and an appraisal of current knowledge. Syst Bot 32:366–378. doi:10.1600/036364407781179653
Bruyns PV, Klak C, Hanacek P (2011) Age and diversity in Old World succulent species of Euphorbia (Euphorbiaceae). Taxon 60:1717–1733
Buining AF, Donald JD (1963) Die Gattung Rebutia K.Schumann. Sukkulentenkunde 7(8):96–107
Buining AF, Donald JD (1965) The revision of the genus Rebutia K.Schumann. Cact Succ J Gr Brit 27:36–41
Carlsen MM, Croat TB (2013) A molecular phylogeny of the species-rich neotropical genus Anthurium (Araceae) based on combined chloroplast and nuclear DNA. Syst Bot 38:576–588. doi:10.1600/036364413X670287
Demaio P, Barfuss MH, Kiesling R, Till W, Chiapella JO (2011) Molecular phylogeny of Gymnocalycium (Cactaceae): assessment of alternative infrageneric systems, a new subgenus, and trends in the evoultion of the genus. Amer J Bot 98:1841–1854. doi:10.3732/ajb.1100054
Echinopseen ZA (1987) Weitere Ergebnisse der Fertilitätsuntersuchungen. Zentrale Arbeitsgruppe Echinopseen—Informationsbrief 10–12:10
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molec Ecol 14:2611–2620. doi:10.1111/j.1365-294X.2005.02553.x
Falush D, Stephens M, Pritchard JK (2007) Inference of population structure using multilocus genotype data: dominant markers and null alleles. Molec Ecol Notes 7:574–578. doi:10.1111/j.1471-8286.2007.01758.x
Farris JS, Kallersjo M, Kluge AG, Bult C (1995) Constructing a significance test for incongruence. Syst Biol 44:570–572. doi:10.2307/2413663
Good-Avila SV, Souza V, Gaut BS, Eguiarte LE (2006) Timing and rate of speciation in Agave (Agavaceae). Proc Natl Acad Sci USA 103:9124–9129. doi:10.1073/pnas.0603312103
Griffith MP, Porter JM (2009) Phylogeny of Opuntioideae (Cactaceae). Int J Pl Sci 170:107–116. doi:10.1086/593048
Guerrero PC, Arroyo MTK, Bustamante RO, Duarte M, Hagemann TK, Walter HE (2011) Phylogenetics and predictive distribution modeling provide insights into the geographic divergence of Eriosyce subgen. Neoporteria (Cactaceae). Pl Syst Evol 297:113–128. doi:10.1007/s00606-011-0512-5
Hernández-Hernández T, Hernández HM, De-Nova JA, Puente R, Eguiarte LE, Magallón S (2011) Phylogenetic relationships and evolution of growth form in Cactaceae (Caryophyllales, Eucotyledoneae). Amer J Bot 98:44–61. doi:10.3732/ajb.1000129
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogeny. Bioinformatics 17:754–755. doi:10.1093/bioinformatics/17.8.754
Hughes C, Eastwood R (2006) Island radiation on a continental scale: exceptional rates of plant diversification after uplift of the Andes. Proc Natl Acad Sci USA 103:10334–10339. doi:10.1073/pnas.0601928103
Hunt D (1999) CITES Cactaceae checklist. Royal Botanic Gardens Kew, Richmond
Hunt D (2006) The new cactus lexicon. dh Books, Milborn Port
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Molec Biol Evol 23:254–267. doi:10.1093/molbev/msj030
Ingvarsson PK, Ribstein S, Taylor DR (2003) Molecular evolution of insertions and deletion in the chloroplast genome of Silene. Molec Biol Evol 20:1737–1740. doi:10.1093/molbev/msg163
Klak C, Reeves G, Hedderson T (2004) Unmatched tempo of evolution in semi-desert ice plants. Nature 427:63-65. doi:10.1038/nature02243
Koopman WJM, Wissemann V, De Cock K, Van Huylenbroeck J, De Riek J, Sabatino GJH, Visser D, Vosman B, Ritz CM, Maes B, Werlemark G, Nybom H, Debener T, Linde M, Smulders MJA (2008) AFLP markers as a tool to reconstruct complex relationships: a case study in Rosa (Rosaceae). Amer J Bot 95:353–366. doi:10.3732/ajb.95.3.353
Krainz H (1967) Die Kakteen: eine Gesamtdarstellung der eingeführten Arten nebst Anzucht- und Pflegeanweisungen. Franckh, Stuttgart
Mosti S, Bandara NL, Papini A (2011a) Further insights and new combinations in Aylostera (Cactaceae) based on molecular and morphological data. Pakistan J Bot 43:2769–2785
Mosti S, Fiorini G, Papini A (2011b) Karyological investigations on several species of the genus Rebutia sect. Digitorebutia (Cactaceae). Caryologia 64:350–359
Mutke J, Barthlott W (2005) Patterns of vascular plant diversity at continental to global scale. Biol Skr 55:521–531
Myers N, Mittermeier RA, Mittermeier CG, da Fonseca GAB, Kent J (2000) Biodiversity hotspots for conservation priorities. Nature 403:853–858. doi:10.1038/35002501
Nyffeler R (2002) Phylogenetic relationships in the cactus family (Cactaceae) based on evidence from trnK/matK and trnL-trnF sequences. Amer J Bot 89:312–326. doi:10.3732/ajb.89.2.312
Nyffeler R, Eggli U (2010) A farewell to dated ideas and concepts: molecular phylogenetics and a revised suprageneric classification of the family Cactaceae. Schumannia 6:109–149. doi:10.5167/uzh-43285
Nylander JAA (2004) MrModeltest. Evolutionary Biology Centre, Uppsala University, Uppsala, Program distributed by the author. https://github.com/nylander/MrModeltest2. Accessed 8 Dec 2014
Pennigton RT, Lewis GP, Ratter JA (2006) Neotropical savannas and seasonally dry forests: plant diversity, biogeography, and conservation. CRC Press, London
Pilbeam J (1997) Rebutia. Cirio Publishing Services, Southampton
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Ritz CM, Martins L, Mecklenburg R, Goremykin V, Hellwig FH (2007) The molecular phylogeny of Rebutia (Cactaceae) and its allies demonstrates the influence of paleogeography on the evolution of South American mountain cacti. Amer J Bot 94:1321–1332. doi:10.3732/ajb.94.8.1321
Ritz CM, Reiker J, Charles G, Hoxey P, Hunt D, Lowry M, Stuppy W, Taylor N (2012) Molecular phylogeny and character evolution in terete-stemmed Andean opuntias (Cactaceae-Opuntioideae). Molec Phylogen Evol 65:668–681. doi:10.1016/j.ympev.2012.07.027
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Molec Ecol Notes 4:137–138. doi:10.1046/j.1471-8286.2003.00566.x
Rowley GD (2009) Rebutia reappraised. CactusWorld 27:88–90
Savolainen V, Manen JF, Douzery E, Spichiger R (1994) Molecular phylogeny of families related to Celastrales based on rbcL 5′ flanking sequences. Molec Phylogen Evol 3:27–37. doi:10.1006/mpev.1994.1004
Schlumpberger BO, Renner SS (2012) Molecular phylogenetics of Echinopsis (Cactaceae): polyphyly at all levels and convergent evolution of pollination modes and growth forms. Amer J Bot 99:1335–1349. doi:10.3732/ajb.1100288
Schumann K (1895) Rebutia minuscula, eine neue Gattung der Kakteen. Monatsschr Kakteenk 5:102–105
Shaw J, Lickey EB, Schilling EE, Small RL (2007) Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III. Amer J Bot 94:275–288. doi:10.3732/ajb.94.3.275
Simmons MP (2004) Independence of alignment and tree search. Molec Phylogen Evol 31:874–879. doi:10.1016/j.ympev.2003.10.008
Stamatakis A (2014) RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 30:1312–1313. doi:10.1093/bioinformatics/btu033
Swofford DL (2002) Paup* 4.0. Phylogenetic analyses using parsimony (and other methods). Sinauer Associates, Sunderland, Massachusetts
Vos P, Hogers R, Bleeker M, Reijans M, Van de Lee T, Hornes M, Fritjers A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucl Acids Res 23:4407–4414. doi:10.1093/nar/23.21.4407
Weising K, Nybom H, Wolff K, Kahl G (2005) DNA fingerprinting in plants—principles, methods and applications. Taylor & Francis Group, Abingdon. doi:10.1017/S0014479705213339
Wissemann V, Ritz CM (2005) The genus Rosa (Rosoideae, Rosaceae) revisited: molecular analysis of nrITS-1 and atpB-rbcL intergenic spacer (IGS) versus conventional taxonomy. Bot J Linn Soc 147:275–290. doi:10.1111/j.1095-8339.2005.00368.x
Yang ZH, Rannala B (1997) Bayesian phylogenetic inference using DNA sequences: a Markov Chain Monte Carlo method. Molec Biol Evol 14:717–724
Acknowledgments
The authors thank the following: R. Märtin (Jena, Germany), K. Meißner (Dresden, Germany) and U. Eggli (Sukkulentensammlung Zürich, Switzerland, ZSS) for providing plant material; H. Krufzik, E. Schirmeier (Justus Liebig University Gießen, Germany), M. Schwager (Senckenberg Museum of Natural History Görlitz, Germany) and the colleagues of the laboratory at the Biodiversity and Climate Research Centre in Frankfurt/Main (BiK-F) for excellent technical support; V. Wissemann (Justus Liebig University Gießen, Germany) for the opportunity to work in his lab; A. Allspach, S. Dressler and R. Döring (Senckenberg Research Institute and Natural History Museum Frankfurt/Main, Germany) for their help in incorporating the plant samples into the spirit collection of the Herbarium Senckenbergianum Frankfurt (FR). We thank U. Eggli and R. Kiesling (Argentina, Mendoza, CCT-CONICET) for their help with nomenclatural questions and all members of the Studiengemeinschaft Südamerikanische Kakteen e.V. for the financial support of this study and for fruitful discussions on the manuscript, especially checking the key to species based on their own plant material. We thank J. C. Hutchinson, K. Wesche (Senckenberg Museum of Natural History Görlitz, Germany) and the anonymous reviewers for helpful comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling editor: Mark Mort.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Information on Electronic Supplementary Material
Information on Electronic Supplementary Material
Online Resource 1. Complete list of accessions analysed
Online Resource 2. Alignment of the combined cp markers: trnS-trnG IGS and atpB-rbcL IGS
Online Resource 3. AFLP scoring table
Rights and permissions
About this article
Cite this article
Ritz, C.M., Fickenscher, K., Föller, J. et al. Molecular phylogenetic relationships of the Andean genus Aylostera Speg. (Cactaceae, Trichocereeae), a new classification and a morphological identification key. Plant Syst Evol 302, 763–780 (2016). https://doi.org/10.1007/s00606-016-1296-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-016-1296-4