Abstract
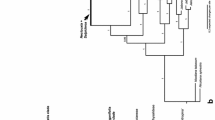
To reconstruct phylogeny and verify the monophyly of major subgroups, a total of 52 species representing almost all species of Salsoleae s.l. in China were sampled, with analysis based on three molecular markers (nrDNA ITS, cpDNA psbB–psbH and rbcL), using maximum parsimony, maximum likelihood, and Bayesian inference methods. Our molecular evidence provides strong support for the following: (1) Camphorosmeae is nested within Salsoleae s.l. instead of the previously suggested sister relationship. (2) Tribe Salsoleae s.l. is monophyletic and is composed of three monophyletic subunits, Caroxyloneae, the Kali clade, and Salsoleae s.str. (3) Climacoptera is separated from Salsola s.l. It does not form a monophyletic group but is split into two monophyletic parts, Climacoptera I and Climacoptera II. (4) Halogeton is clearly polyphyletic, as are Anabasis and the genus Salsola s.l. (5) Caroxylon, Haloxylon, Kali, and Petrosimonia are well-supported monophyletic genera. Additional evidence is needed regarding the monophyly of Halimocnemis, which remains unclear.




Similar content being viewed by others
References
Akhani H (2004) Halophytic vegetation of Iran: towards a syntaxonomical classification. Ann Bot (Rome) 4:66–82
Akhani H, Trimborn P, Ziegler H (1997) Photosynthetic pathways in Chenopodiaceae from Africa, Asia and Europe with their ecological, phytogeographical and taxonomical importance. Plant Syst Evol 206:187–221
Akhani H, Ghobadnejhad M, Hashemi SM (2003) Ecology, biogeography and pollen morphology of Bienertia cycloptera Bunge ex Boiss. (Chenopodiaceae), an enigmatic C4 plant without Kranz anatomy. Plant Biol 5:167–178
Akhani H, Edwards G, Roalson EH (2007) Diversification of the old world Salsoleae s.l. (Chenopodiaceae): molecular phylogenetic analysis of nuclear and chloroplast data sets and a revised classification. Int J Plant Sci 168:931–956
Assadi M (2001) Chenopodiaceae. In: Assadi M, Khatamsaz M, Maassoumi AA (eds) Flora of Iran, vol 38. Research Institute of Forests and Rangelands, Tehran, pp 27–65
Blackwell WH Jr (1977) The subfamilies of the Chenopodiaceae. Taxon 26:395–397
Borger CP, Yan GJ, Scott JK, Walsh MJ (2008) Salsola tragus or S. australis (Chenopodiaceae) in Australia—untangling taxonomic confusion through molecular and cytological analyses. Aust J Bot 56:600–608
Botschantzev VP (1956) Sbornik rabot po geobotanike, lesovedeniju, paleogeografii floristike: dva novykh roda iz semeistva marevykh. In: Akademiku VN, Sukachevu K (eds) Akademia Nauk SSSR. Izdatel’stvo Akademia Nauk SSSR, Moscow, pp 108–118
Botschantzev VP (1969) The genus Salsola: a concise history of its development and dispersal (in Russian). Bot Zhurn 54:989–1001
Botschantzev VP (1974) Species subsections Caroxylon sections Caroxylon (Thunb.) Fenzl generis Salsola L. (in Russian). Nov Sist Vyssh Rast 11:110–174
Botschantzev VP (1976) Conspectus speciorum sections Coccosalsola Fenzl generis Salsola L. (in Russian). Nov Sist Vyssh Rast 13:74–102
Butnik AA (1979) Types of development of seedlings of Chenopodiaceae Vent. (in Russian). Bot Zhurn 64:834–842 (in Russian)
Cabrera JF, Jacobs SWL, Kadereit G (2009) Phylogeny of the Australian Camphorosmeae (Chenopodiaceae) and the taxonomic significance of the fruiting perianth. Int J Plant Sci 170:505–521
Casati P, Andreo CS, Edwards GE (1999) Characterization of NADP-malic enzyme from two species of Chenopodiaceae: Haloxylon persicum (C4) and Chenopodium album (C3). Phytochemistry 52:985–992
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Freitag H (1997) Salsola L. (Chenopodiaceae). In: Rechinger KH (ed) Flora Iranica, vol 172. Akademische Druck und Verlagsanstalt, Graz, pp 154–255
Fu LK, Zhang XC, Qin HN, Ma JS (1993) Index herbariorum sinicorum (in Chinese). Chinese Science and Technology Press, Beijing, pp 425–457
Grubov VI (1999) Chenopodiaceae. In: Plants of Central Asia, vol 2. Science Publishers, Enfield, pp 87–133
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Hillis DM, Bull JJ (1993) An empirical test of bootstrapping as a method for assessing confidence in phylogenetic analysis. Syst Biol 42:182–192
Holmgren PK, Holmgren NH (1998) (continuously updated) Index herbariorum. http://sciweb.nybg.org/science2/IndexHerbariorum.asp
Huelsenbeck JP, Rannala B (2004) Frequentist properties of Bayesian posterior probabilities of phylogenetic trees under simple and complex substitution models. Syst Biol 53:904–913
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Iljin MM (1936) Chenopodiaceae. In: Siskin BK (ed) Flora SSSR, vol 6 (in Russian). Izdatel’stvo Akademii Nauk SSSR, Leningrad, pp 2–354
Johnson LA, Soltis DE (1995) Phylogenetic inference in Saxifragaceae sensu stricto and Gilia (Polemoniaceae) using matK sequences. Ann Mo Bot Gard 82:149–175
Kadereit G, Borsch T, Weising K, Freitag H (2003) Phylogeny of Amaranthaceae and Chenopodiaceae and the evolution of C4 photosynthesis. Int J Plant Sci 164:959–986
Kadereit G, Gotzek D, Jacobs S, Freitag H (2005) Origin and age of Australian Chenopodiaceae. Org Divers Evol 5:59–80
Kang Y, Zhang ML, Chen ZD (2003) A preliminary phylogenetic study of the subgenus Pogonophace (Astragalus) in China based on ITS sequence data. Acta Bot Sin 45:140–145
Kapralov MV, Akhani H, Voznesenskaya EV, Edwards G, Franceschi V, Roalson EH (2006) Phylogenetic relationships in the Salicornioideae/Suaedoideae/Salsoloideae s.l. (Chenopodiaceae) clade and a clarification of the phylogenetic position of Bienertia and Alexandra using multiple DNA sequence datasets. Syst Bot 31:571–585
Kühn U, Bittrich V, Carolin R, Freitag H, Hedge IC, Uotila P, Wilson PG (1993) Chenopodiaceae. In: Kubitzki K, Rohwer JG, Bittrich V (eds) The families and genera of vascular plants, vol 2. Springer, Berlin, pp 253–281
Liu YX (1995) Observations on the formation of Chinese desert floras (in Chinese with English abstract). Acta Phytotax Sin 33:131–143
Meyer CA (1829) Generae Chenopodearum. In: Ledebour CF (ed) Flora Altaica, vol 2. Reimer, Berlin, pp 370–371
Moquin-Tandon A (1840) Chenopodearum monographica enumeratio. Loss, Paris, p 182
Moquin-Tandon A (1849) Salsolaceae. In: de Candolle AP (ed) Prodromus systematis naturalis regni vegetabilis, vol 13. Masson, Paris, pp 41–219
Posada D, Crandall KA (1998) Modeltest: testing the model of DNA substitution. Bioinformatics 14:817–818
Pyankov VI, Voznesenskaya EV, Kuz’min AN, Ku MSB, Ganko E, Franceschi VR, Black CC, Edwards GE (2000) Occurrence of C3 and C4 photosynthesis in cotyledons and leaves of Salsola species (Chenopodiaceae). Photosynth Res 63:69–84
Pyankov VI, Artyusheva EG, Edwards GE, Black CC, Soltis PS (2001a) Phylogenetic analysis of tribe Salsoleae (Chenopodiaceae) based on ribosomal ITS sequences: implications for the evolution of photosynthesis types. Am J Bot 88:1189–1198
Pyankov VI, Ziegler H, Kuz’min A, Edwards G (2001b) Origin and evolution of C4 photosynthesis in the tribe Salsoleae (Chenopodiaceae) based on anatomical and biochemical types in leaves and cotyledons. Plant Syst Evol 230:43–74
Rilke S (1999) Species diversity and polymorphism in Salsola sect. Salsola sensu lato (Chenopodiacaeae). Syst Geogr Pl 68:305–314
Schütze P, Freitag H, Weising K (2003) An integrated molecular and morphological study of the subfamily Suaedoideae Ulbr. (Chenopodiaceae). Plant Syst Evol 239:257–286
Sukhorukov AP (2008) Fruit anatomy of the genus Anabasis (Salsoloideae, Chenopodiaceae). Aust Syst Bot 21:431–442
Swofford DL (2002) PAUP*: phylogenetic analysis using parsimony (* and other methods), version 4.0. Sinauer, Sunderland
Takhtajan A (2009) Flowering plants, vol 1, 2nd edn. Springer, Berlin
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24:4876–4882
Tzvelev NN (1993) Notes on Chenopodiaceae of Eastern Europe. Ukr Bot Zhurn 50:78–85
Ulbrich E (1934) Chenopodiaceae. In: Engler A, Prantl K (eds) Die natürlichen Pflanzenfamilien, 2nd edn. Duncker & Humblot, Leipzig, pp 379–584
Voznesenskaya EV (1976) The ultrastructure of assimilating organs of some species of the family Chenopodiaceae, II (in Russian). Bot Zhurn 61:1546–1557
Voznesenskaya EV, Artyusheva EG, Franceschi VR, Pyankov VI, Kiirats O, Ku MSB, Edwards GE (2001) Salsola arbusculiformis, a C3–C4 intermediate in Salsoleae (Chenopodiaceae). Ann Bot 88:337–348
Wang RZ (2007) C4 plants in the deserts of China: occurrence of C4 photosynthesis and its morphological functional types. Photosynthetica 45:167–171
Wei Y, Dong M, Huang ZY, Tan DY (2008) Factors influencing seed germination of Salsola affinis (Chenopodiaceae), a dominant annual halophyte inhabiting the deserts of Xinjiang, China. Flora 203:134–140
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis M, Gelfand D, Sninsky J, White T (eds) PCR protocols: a guide to methods and applications. Academic, San Diego, pp 315–322
Williams JT, Ford-Lloyd BV (1974) The systematics of the Chenopodiaceae. Taxon 23:353–354
Wilson PG (1984) Chenopodiaceae. In: George AS (ed) Flora of Australia, vol 4. Australian Government Publishing Service, Canberra, pp 313–317
Xu DH, Abe J, Sakai M, Kanazawa A, Shimamoto Y (2000) Sequence variation of non-coding regions of chloroplast DNA of soybean and related wild species and its implications for the evolution of different chloroplast haplotypes. Theor Appl Genet 101:724–732
Zhao KF, Fan H, Ungar IA (2002) Survey of halophyte species in China. Plant Sci 163:491–498
Zhu GL (1996) Origin, differentiation, and geographic distribution of the Chenopodiaceae (in Chinese with English abstract). Acta Phytotax Sin 34:486–504
Zhu GL, Mosyankin SL, Clemants SE (2003) Chenopodiaceae. In: Wu ZY, Raven PH (eds) Flora of China, vol 5. Science Press, Beijing, pp 354–414
Zurawski G, Perrot B, Bottomley W, Whitfeld PR (1981) The structure of the gene for the large subunit of ribulose 1, 5-bisphosphate carboxylase from spinach chloroplast DNA. Nucleic Acids Res 9:3251–3270
Acknowledgments
Thanks to Prof. P. Yan for providing Salsoleae s.l. field collections from Xinjiang Province, China, Dr. D. M. Williams (London, UK) for helpful comments on the manuscript, Mrs. Lorraine Williams (London, UK) for improving the English of the manuscript, and two anonymous reviewers for valuable comments on a previous version. This research was funded by the National Basic Research Program of China (2009CB825104), Xinjiang Institute of Ecology and Geography, Chinese Academy of Sciences.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wen, ZB., Zhang, ML., Zhu, GL. et al. Phylogeny of Salsoleae s.l. (Chenopodiaceae) based on DNA sequence data from ITS, psbB–psbH, and rbcL, with emphasis on taxa of northwestern China. Plant Syst Evol 288, 25–42 (2010). https://doi.org/10.1007/s00606-010-0310-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-010-0310-5